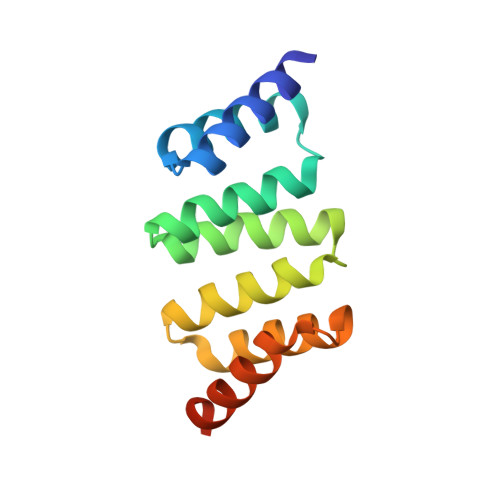
Tailored design of protein nanoparticle scaffolds for multivalent presentation of viral glycoprotein antigens.
Ueda, G., Antanasijevic, A., Fallas, J.A., Sheffler, W., Copps, J., Ellis, D., Hutchinson, G.B., Moyer, A., Yasmeen, A., Tsybovsky, Y., Park, Y.J., Bick, M.J., Sankaran, B., Gillespie, R.A., Brouwer, P.J., Zwart, P.H., Veesler, D., Kanekiyo, M., Graham, B.S., Sanders, R.W., Moore, J.P., Klasse, P.J., Ward, A.B., King, N.P., Baker, D.(2020) Elife 9
- PubMed: 32748788 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.57659
- Primary Citation Related Structures:
6V8E, 6VEH, 6VFH, 6VFI, 6VFJ - PubMed Abstract:
Multivalent presentation of viral glycoproteins can substantially increase the elicitation of antigen-specific antibodies. To enable a new generation of anti-viral vaccines, we designed self-assembling protein nanoparticles with geometries tailored to present the ectodomains of influenza, HIV, and RSV viral glycoprotein trimers. We first de novo designed trimers tailored for antigen fusion, featuring N-terminal helices positioned to match the C termini of the viral glycoproteins. Trimers that experimentally adopted their designed configurations were incorporated as components of tetrahedral, octahedral, and icosahedral nanoparticles, which were characterized by cryo-electron microscopy and assessed for their ability to present viral glycoproteins. Electron microscopy and antibody binding experiments demonstrated that the designed nanoparticles presented antigenically intact prefusion HIV-1 Env, influenza hemagglutinin, and RSV F trimers in the predicted geometries. This work demonstrates that antigen-displaying protein nanoparticles can be designed from scratch, and provides a systematic way to investigate the influence of antigen presentation geometry on the immune response to vaccination.
- Department of Biochemistry, University of Washington, Seattle, United States.
Organizational Affiliation: