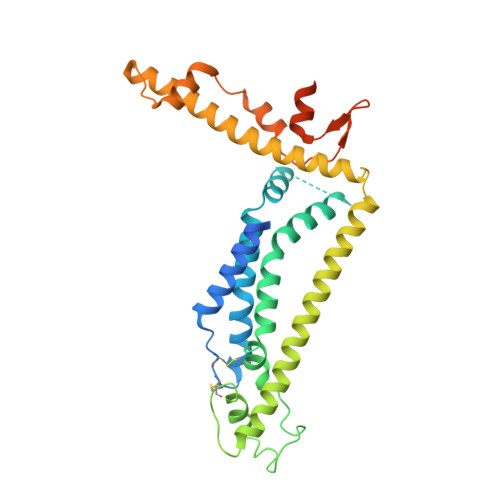
Structure and assembly of calcium homeostasis modulator proteins.
Syrjanen, J.L., Michalski, K., Chou, T.H., Grant, T., Rao, S., Simorowski, N., Tucker, S.J., Grigorieff, N., Furukawa, H.(2020) Nat Struct Mol Biol 27: 150-159
- PubMed: 31988524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0369-9
- Primary Citation Related Structures:
6VAI, 6VAK, 6VAL, 6VAM - PubMed Abstract:
The biological membranes of many cell types contain large-pore channels through which a wide variety of ions and metabolites permeate. Examples include connexin, innexin and pannexin, which form gap junctions and/or bona fide cell surface channels. The most recently identified large-pore channels are the calcium homeostasis modulators (CALHMs), through which ions and ATP permeate in a voltage-dependent manner to control neuronal excitability, taste signaling and pathologies of depression and Alzheimer's disease. Despite such critical biological roles, the structures and patterns of their oligomeric assembly remain unclear. Here, we reveal the structures of two CALHMs, chicken CALHM1 and human CALHM2, by single-particle cryo-electron microscopy (cryo-EM), which show novel assembly of the four transmembrane helices into channels of octamers and undecamers, respectively. Furthermore, molecular dynamics simulations suggest that lipids can favorably assemble into a bilayer within the larger CALHM2 pore, but not within CALHM1, demonstrating the potential correlation between pore size, lipid accommodation and channel activity.
- WM Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, USA.
Organizational Affiliation: