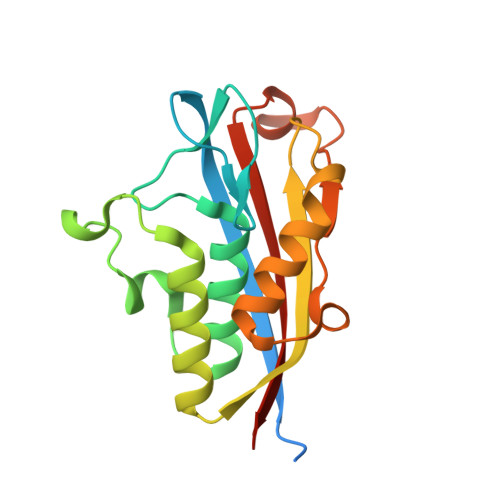
Analysis of Burkholderia pseudomallei IspF in complex with sulfapyridine, sulfamonomethoxine, ethoxzolamide and acetazolamide.
Grote, D., Stewart, C.G., Daraji, D.G., Enayati, P., Braverman, K.N., Romanaggi, C., Bolejack, M.J., Yano, J.K., Abendroth, J., Dranow, D.M., Pierce, P.G., Lorimer, D.D., Horanyi, P.S., Staker, B.L., Edwards, T.E., Myler, P.J., Horn, J.R., Hagen, T.J.(2025) Acta Crystallogr F Struct Biol Commun 81: 138-145
- PubMed: 40035494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25001414
- Primary Citation Related Structures:
6V3M - PubMed Abstract:
The methylerythritol phosphate (MEP) pathway is a metabolic pathway that produces the isoprenoids isopentyl pyrophosphate and dimethylallyl pyrophosphate. Notably, the MEP pathway is present in bacteria and not in mammals, which makes the enzymes of the MEP pathway attractive targets for the discovery of new anti-infective agents due to the reduced chances of off-target interactions leading to side effects. There are seven enzymes in the MEP pathway, the fifth of which is IspF. Crystal structures of Burkholderia pseudomallei IspF were determined with five different sulfonamide ligands bound. The sulfonamide-containing ligands were ethoxzolamide, acetazolamide, sulfapyridine and sulfamonomethoxine. The fifth bound ligand was a synthetic analog of acetazolamide. All ligands coordinated to the active-site Zn +2 ion through the sulfonamide group, although sulfapyridine and sulfamonomethoxine, both of which are known antibacterial agents, possess similar binding interactions that are distinct from the other three sulfonamides. These structural data will aid in the discovery of new IspF inhibitors.
- Department of Chemistry and Biochemistry, Northern Illinois University, 1425 Lincoln Highway, DeKalb, IL 60115, USA.
Organizational Affiliation: