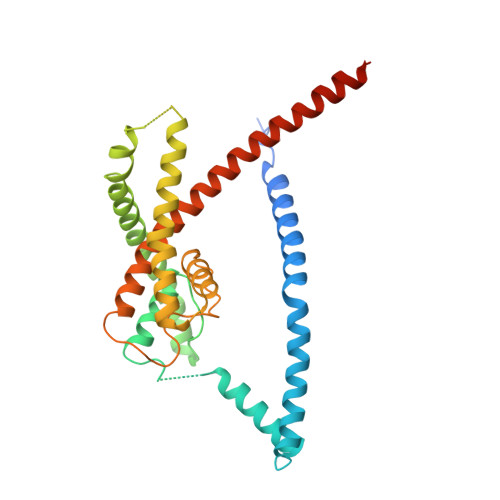
Polynuclear Ruthenium Amines Inhibit K2PChannels via a "Finger in the Dam" Mechanism.
Pope, L., Lolicato, M., Minor Jr., D.L.(2020) Cell Chem Biol
- PubMed: 32059793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2020.01.011
- Primary Citation Related Structures:
6V36, 6V37, 6V3C, 6V3I - PubMed Abstract:
The trinuclear ruthenium amine ruthenium red (RuR) inhibits diverse ion channels, including K 2P potassium channels, TRPs, the calcium uniporter, CALHMs, ryanodine receptors, and Piezos. Despite this extraordinary array, there is limited information for how RuR engages targets. Here, using X-ray crystallographic and electrophysiological studies of an RuR-sensitive K 2P , K 2P 2.1 (TREK-1) I110D, we show that RuR acts by binding an acidic residue pair comprising the "Keystone inhibitor site" under the K 2P CAP domain archway above the channel pore. We further establish that Ru360, a dinuclear ruthenium amine not known to affect K 2P s, inhibits RuR-sensitive K 2P s using the same mechanism. Structural knowledge enabled a generalizable design strategy for creating K 2P RuR "super-responders" having nanomolar sensitivity. Together, the data define a "finger in the dam" inhibition mechanism acting at a novel K 2P inhibitor binding site. These findings highlight the polysite nature of K 2P pharmacology and provide a new framework for K 2P inhibitor development.
- Cardiovascular Research Institute, University of California, San Francisco, CA 93858-2330, USA.
Organizational Affiliation: