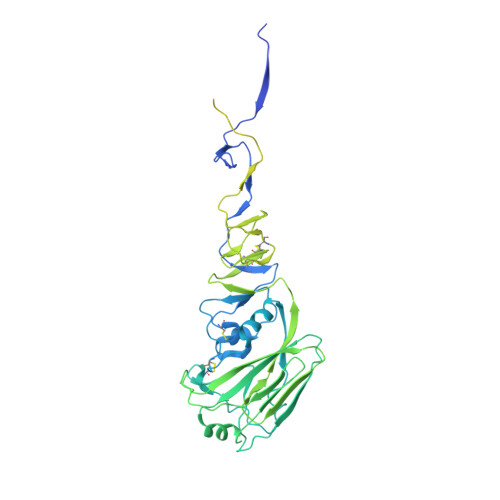
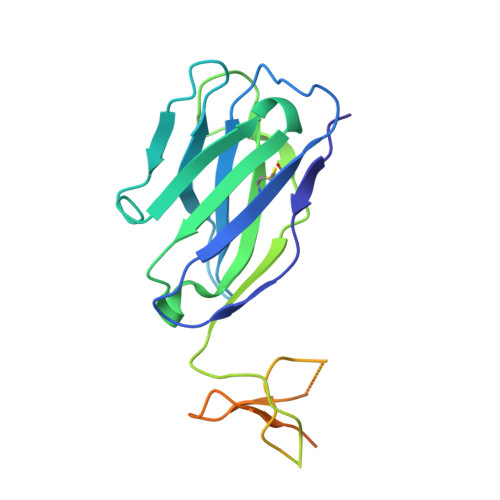
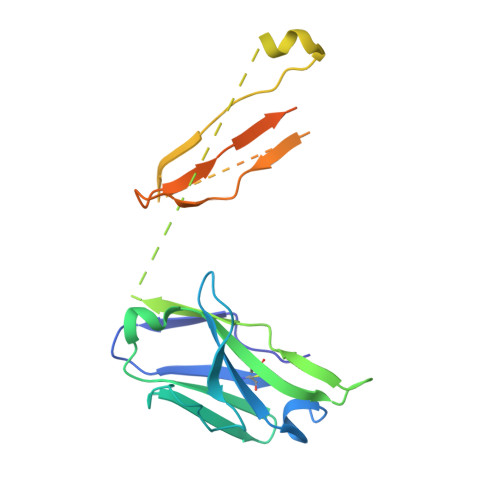
Crystal structure of influenza A virus hemagglutinin from A/Ohio/09/2015 bound to the stalk-binding CR6261 antibody Fab
Edwards, T.E., Fox III, D., Horanyi, P.S., Lorimer, D.D., Calhoun, B., Conrady, D.G., Abendroth, J., Seattle Structural Genomics Center for Infectious Disease (SSGCID)To be published.