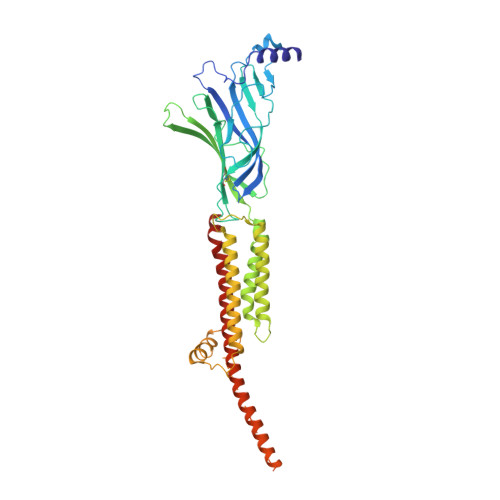
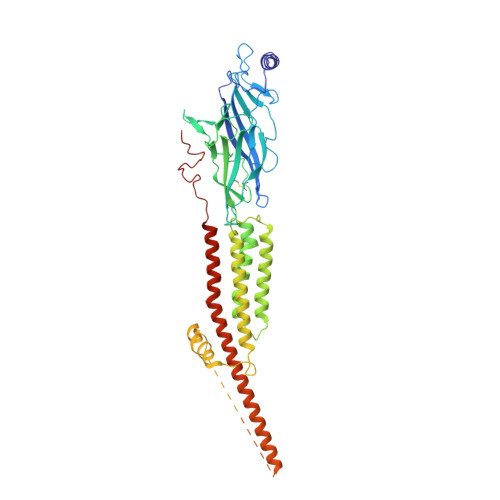
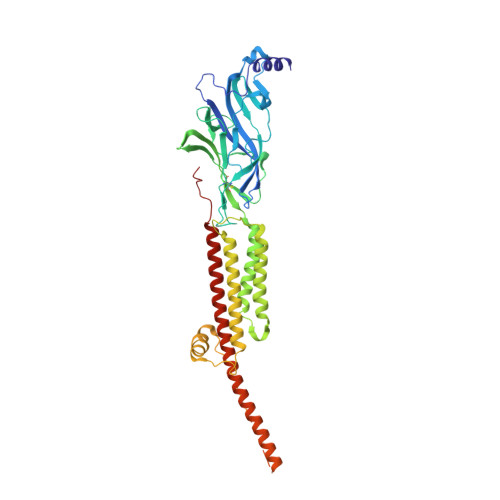
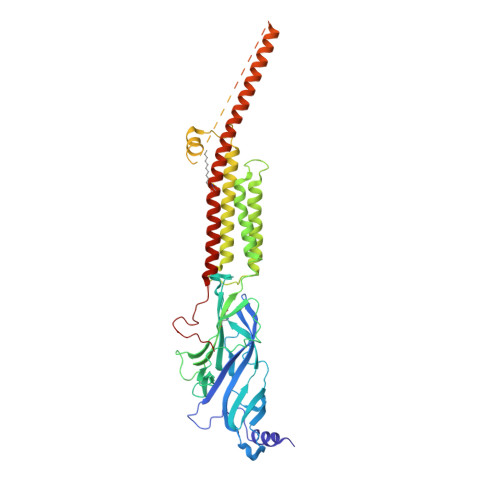
Structure of the Native Muscle-type Nicotinic Receptor and Inhibition by Snake Venom Toxins.
Rahman, M.M., Teng, J., Worrell, B.T., Noviello, C.M., Lee, M., Karlin, A., Stowell, M.H.B., Hibbs, R.E.(2020) Neuron 106: 952-962.e5
- PubMed: 32275860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.neuron.2020.03.012
- Primary Citation Related Structures:
6UWZ - PubMed Abstract:
The nicotinic acetylcholine receptor, a pentameric ligand-gated ion channel, converts the free energy of binding of the neurotransmitter acetylcholine into opening of its central pore. Here we present the first high-resolution structure of the receptor type found in muscle-endplate membrane and in the muscle-derived electric tissues of fish. The native receptor was purified from Torpedo electric tissue and functionally reconstituted in lipids optimal for cryo-electron microscopy. The receptor was stabilized in a closed state by the binding of α-bungarotoxin. The structure reveals the binding of a toxin molecule at each of two subunit interfaces in a manner that would block the binding of acetylcholine. It also reveals a closed gate in the ion-conducting pore, formed by hydrophobic amino acid side chains, located ∼60 Å from the toxin binding sites. The structure provides a framework for understanding gating in ligand-gated channels and how mutations in the acetylcholine receptor cause congenital myasthenic syndromes.
- Department of Neuroscience, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: