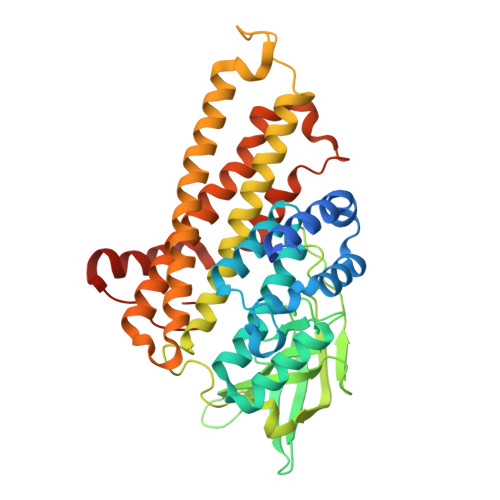
Structure and function of the two-component flavin-dependent methanesulfinate monooxygenase within bacterial sulfur assimilation.
Soule, J., Gnann, A.D., Gonzalez, R., Parker, M.J., McKenna, K.C., Nguyen, S.V., Phan, N.T., Wicht, D.K., Dowling, D.P.(2020) Biochem Biophys Res Commun 522: 107-112
- PubMed: 31753487 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.11.008
- Primary Citation Related Structures:
6UUG - PubMed Abstract:
Methyl sulfur compounds are a rich source of environmental sulfur for microorganisms, but their use requires redox systems. The bacterial sfn and msu operons contain two-component flavin-dependent monooxygenases for dimethylsulfone (DMSO 2 ) assimilation: SfnG converts DMSO 2 to methanesulfinate (MSI - ), and MsuD converts methanesulfonate (MS - ) to sulfite. However, the enzymatic oxidation of MSI - to MS - has not been demonstrated, and the function of the last enzyme of the msu operon (MsuC) is unresolved. We employed crystallographic and biochemical studies to identify the function of MsuC from Pseudomonas fluorescens. The crystal structure of MsuC adopts the acyl-CoA dehydrogenase fold with putative binding sites for flavin and MSI - , and functional assays of MsuC in the presence of its oxidoreductase MsuE, FMN, and NADH confirm the enzymatic generation of MS - . These studies reveal that MsuC converts MSI - to MS - in sulfite biosynthesis from DMSO 2 .
- Department of Chemistry, University of Massachusetts Boston, Boston, MA, 02125, USA.
Organizational Affiliation: