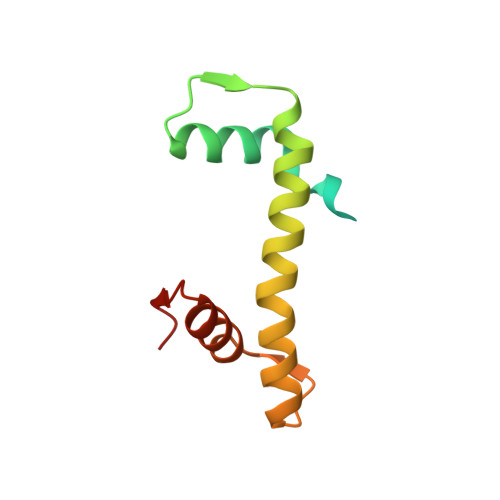
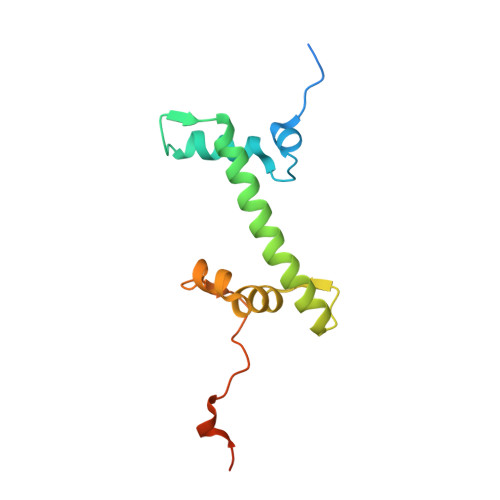
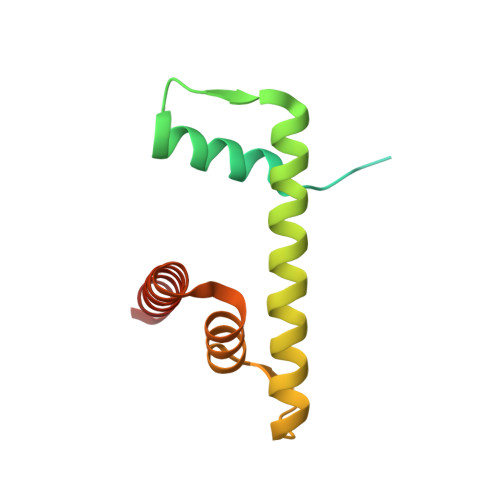
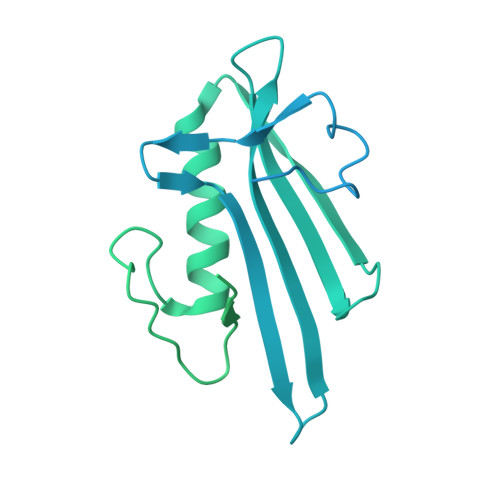
Bridging of nucleosome-proximal DNA double-strand breaks by PARP2 enhances its interaction with HPF1.
Gaullier, G., Roberts, G., Muthurajan, U.M., Bowerman, S., Rudolph, J., Mahadevan, J., Jha, A., Rae, P.S., Luger, K.(2020) PLoS One 15: e0240932-e0240932
- PubMed: 33141820 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0240932
- Primary Citation Related Structures:
6USJ - PubMed Abstract:
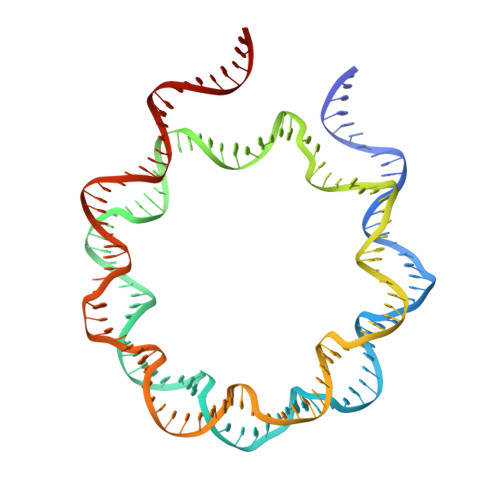
Poly(ADP-ribose) Polymerase 2 (PARP2) is one of three DNA-dependent PARPs involved in the detection of DNA damage. Upon binding to DNA double-strand breaks, PARP2 uses nicotinamide adenine dinucleotide to synthesize poly(ADP-ribose) (PAR) onto itself and other proteins, including histones. PAR chains in turn promote the DNA damage response by recruiting downstream repair factors. These early steps of DNA damage signaling are relevant for understanding how genome integrity is maintained and how their failure leads to genome instability or cancer. There is no structural information on DNA double-strand break detection in the context of chromatin. Here we present a cryo-EM structure of two nucleosomes bridged by human PARP2 and confirm that PARP2 bridges DNA ends in the context of nucleosomes bearing short linker DNA. We demonstrate that the conformation of PARP2 bound to damaged chromatin provides a binding platform for the regulatory protein Histone PARylation Factor 1 (HPF1), and that the resulting HPF1•PARP2•nucleosome complex is enzymatically active. Our results contribute to a structural view of the early steps of the DNA damage response in chromatin.
- Department of Biochemistry, University of Colorado Boulder, Boulder, CO, United States of America.
Organizational Affiliation: