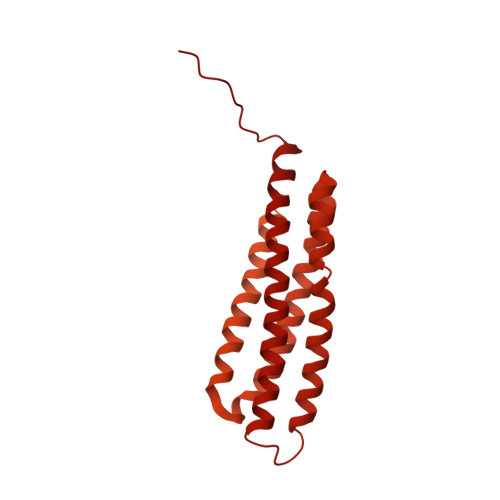
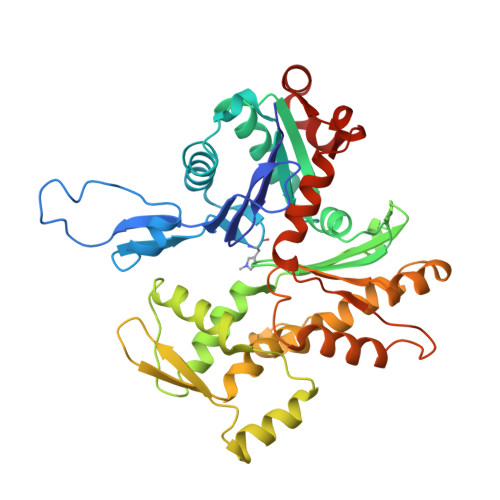
Molecular mechanism for direct actin force-sensing by alpha-catenin.
Mei, L., Espinosa de Los Reyes, S., Reynolds, M.J., Leicher, R., Liu, S., Alushin, G.M.(2020) Elife 9
- PubMed: 32969337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.62514
- Primary Citation Related Structures:
6UPV, 6UPW - PubMed Abstract:
The actin cytoskeleton mediates mechanical coupling between cells and their tissue microenvironments. The architecture and composition of actin networks are modulated by force; however, it is unclear how interactions between actin filaments (F-actin) and associated proteins are mechanically regulated. Here we employ both optical trapping and biochemical reconstitution with myosin motor proteins to show single piconewton forces applied solely to F-actin enhance binding by the human version of the essential cell-cell adhesion protein αE-catenin but not its homolog vinculin. Cryo-electron microscopy structures of both proteins bound to F-actin reveal unique rearrangements that facilitate their flexible C-termini refolding to engage distinct interfaces. Truncating α-catenin's C-terminus eliminates force-activated F-actin binding, and addition of this motif to vinculin confers force-activated binding, demonstrating that α-catenin's C-terminus is a modular detector of F-actin tension. Our studies establish that piconewton force on F-actin can enhance partner binding, which we propose mechanically regulates cellular adhesion through α-catenin.
- Laboratory of Structural Biophysics and Mechanobiology, The Rockefeller University, New York, United States.
Organizational Affiliation: