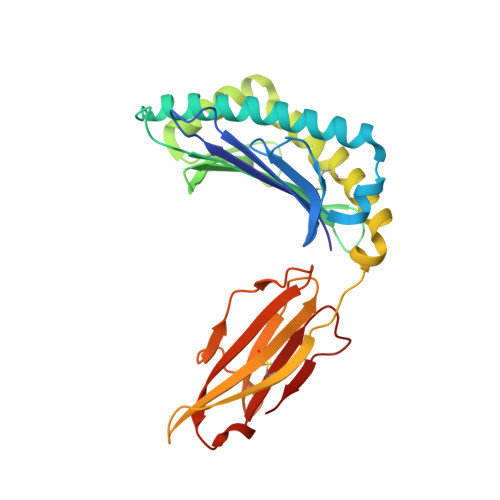
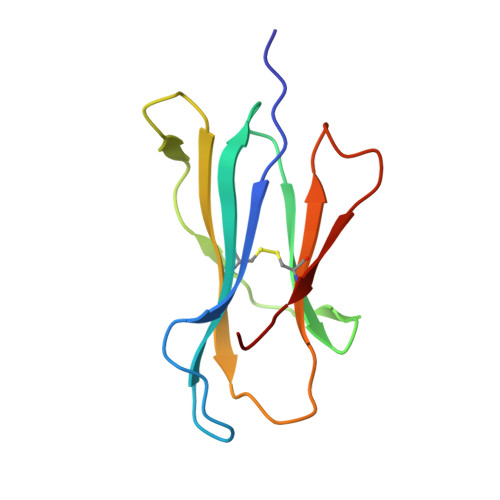
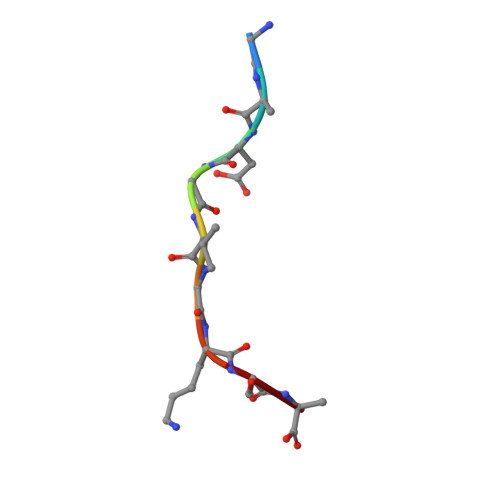
High-affinity oligoclonal TCRs define effective adoptive T cell therapy targeting mutant KRAS-G12D.
Sim, M.J.W., Lu, J., Spencer, M., Hopkins, F., Tran, E., Rosenberg, S.A., Long, E.O., Sun, P.D.(2020) Proc Natl Acad Sci U S A 117: 12826-12835
- PubMed: 32461371 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1921964117
- Primary Citation Related Structures:
6ULI, 6ULK, 6ULN, 6ULR, 6UON - PubMed Abstract:
Complete cancer regression occurs in a subset of patients following adoptive T cell therapy (ACT) of ex vivo expanded tumor-infiltrating lymphocytes (TILs). However, the low success rate presents a great challenge to broader clinical application. To provide insight into TIL-based immunotherapy, we studied a successful case of ACT where regression was observed against tumors carrying the hotspot mutation G12D in the KRAS oncogene. Four T cell receptors (TCRs) made up the TIL infusion and recognized two KRAS-G12D neoantigens, a nonamer and a decamer, all restricted by human leukocyte antigen (HLA) C*08:02. Three of them (TCR9a, 9b, and 9c) were nonamer-specific, while one was decamer-specific (TCR10). We show that only mutant G12D but not the wild-type peptides stabilized HLA-C*08:02 due to the formation of a critical anchor salt bridge to HLA-C. Therapeutic TCRs exhibited high affinities, ranging from nanomolar to low micromolar. Intriguingly, TCR binding affinities to HLA-C inversely correlated with their persistence in vivo, suggesting the importance of antigenic affinity in the function of therapeutic T cells. Crystal structures of TCR-HLA-C complexes revealed that TCR9a to 9c recognized G12D nonamer with multiple conserved contacts through shared CDR2β and CDR3α. This allowed CDR3β variation to confer different affinities via a variable HLA-C contact, generating an oligoclonal response. TCR10 recognized an induced and distinct G12D decamer conformation. Thus, this successful case of ACT included oligoclonal TCRs of high affinity recognizing distinct conformations of neoantigens. Our study revealed the potential of a structural approach to inform clinical efforts in targeting KRAS-G12D tumors by immunotherapy and has general implications for T cell-based immunotherapies.
- Structural Immunology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases (NIAID), NIH, Rockville, MD 20852.
Organizational Affiliation: