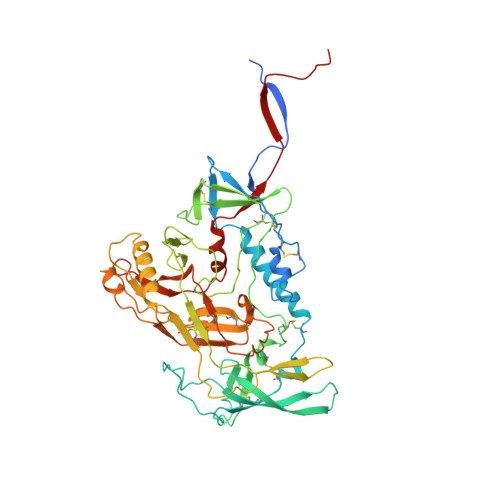
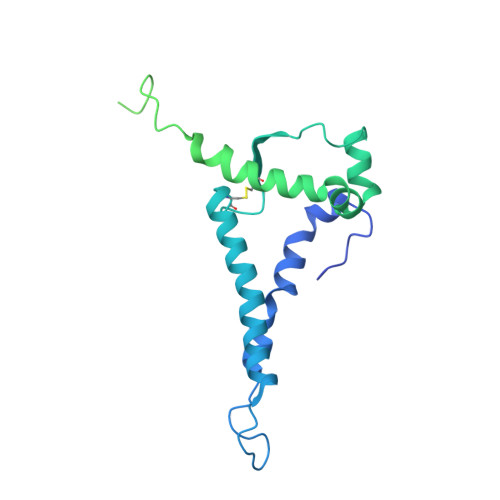
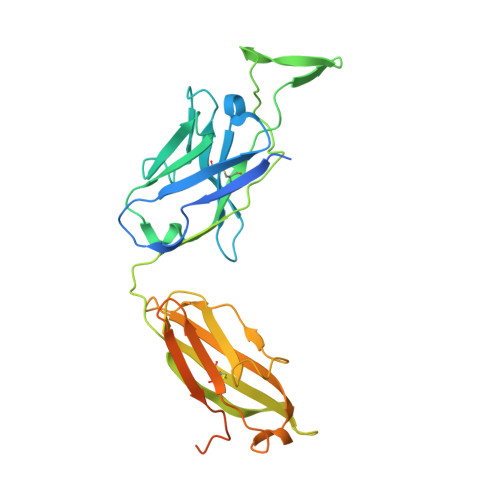
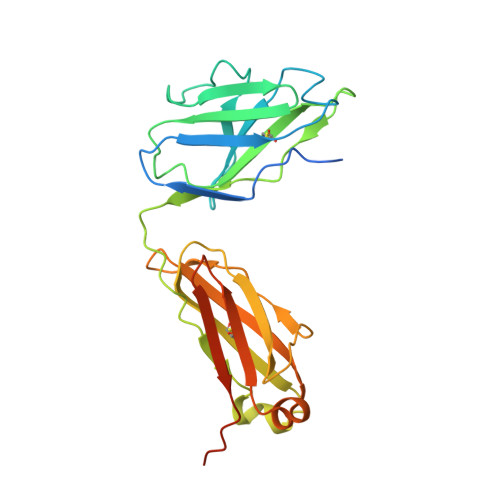
Cryo-EM Structure of Full-length HIV-1 Env Bound With the Fab of Antibody PG16.
Pan, J., Peng, H., Chen, B., Harrison, S.C.(2020) J Mol Biology 432: 1158-1168
- PubMed: 31931014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.11.028
- Primary Citation Related Structures:
6PWU, 6ULC - PubMed Abstract:
The HIV-1 envelope protein (Env) is the target of neutralizing antibodies and the template for vaccine immunogen design. The dynamic conformational equilibrium of trimeric Env influences its antigenicity and potential immunogenicity. Antibodies that bind at the trimer apex stabilize a "closed" conformation characteristic of the most difficult to neutralize isolates. A goal of vaccine development is therefore to mimic the closed conformation in a designed immunogen. A disulfide-stabilized, trimeric Env ectodomain-the "SOSIP" construct-has many of the relevant properties; it is also particularly suitable for structure determination. Some single-molecule studies have, however, suggested that the SOSIP trimer is not a good representation of Env on the surface of a virion or an infected cell. We isolated Env (fully cleaved to gp120 and gp41) from the surface of expressing cells using tagged, apex-binding Fab PG16 and determined the structure of the PG16-Env complex by cryo-EM to an overall resolution of 4.6 Å. Placing the only purification tag on the Fab ensured that the isolated Env was continuously stabilized in its closed, native conformation. The Env structure in this complex corresponds closely to the SOSIP structures determined by both x-ray crystallography and cryo-EM. Although the membrane-interacting elements are not resolved in our reconstruction, we can make inferences about the connection between ectodomain and membrane-proximal external region (MPER) by reference to the published cryo-tomography structure of an Env "spike" and the NMR structure of the MPER-transmembrane segment. We discuss these results in view of the conflicting interpretations in the literature.
- Laboratory of Molecular Medicine, Boston Children's Hospital, 3 Blackfan Circle, Boston, MA, 02115, USA.
Organizational Affiliation: