The terminal sialic acid of stage-specific embryonic antigen-4 has a crucial role in binding to a cancer-targeting antibody.
Soliman, C., Chua, J.X., Vankemmelbeke, M., McIntosh, R.S., Guy, A.J., Spendlove, I., Durrant, L.G., Ramsland, P.A.(2020) J Biological Chem 295: 1009-1020
- PubMed: 31831622 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.011518
- Primary Citation Related Structures:
6UG7, 6UG8, 6UG9, 6UGA - PubMed Abstract:
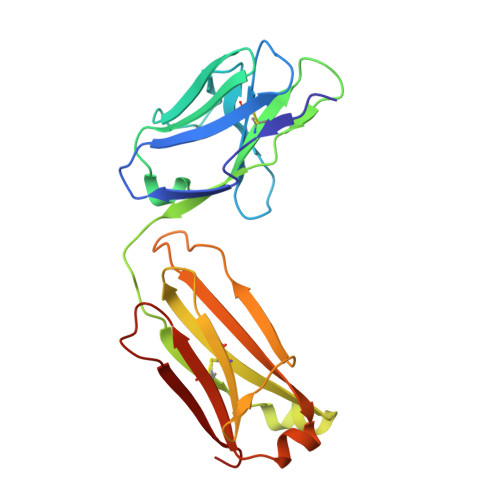
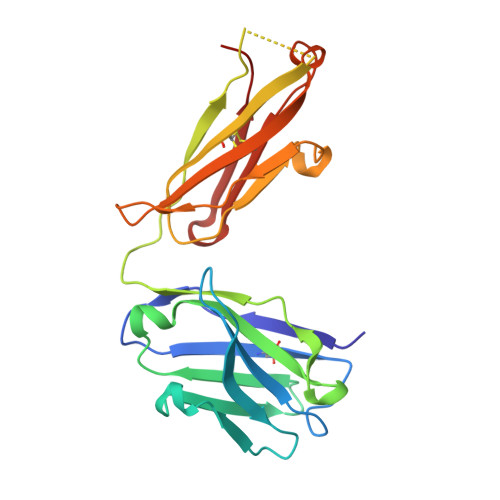
Cancer remains a leading cause of morbidity and mortality worldwide, requiring ongoing development of targeted therapeutics such as monoclonal antibodies. Carbohydrates on embryonic cells are often highly expressed in cancer and are therefore attractive targets for antibodies. Stage-specific embryonic antigen-4 (SSEA-4) is one such glycolipid target expressed in many cancers, including breast and ovarian carcinomas. Here, we defined the structural basis for recognition of SSEA-4 by a novel monospecific chimeric antibody (ch28/11). Five X-ray structures of ch28/11 Fab complexes with the SSEA-4 glycan headgroup, determined at 1.5-2.7 Å resolutions, displayed highly similar three-dimensional structures indicating a stable binding mode. The structures also revealed that by adopting a horseshoe-shaped conformation in a deep groove, the glycan headgroup likely sits flat against the membrane to allow the antibody to interact with SSEA-4 on cancer cells. Moreover, we found that the terminal sialic acid of SSEA-4 plays a dominant role in dictating the exquisite specificity of the ch28/11 antibody. This observation was further supported by molecular dynamics simulations of the ch28/11-glycan complex, which show that SSEA-4 is stabilized by its terminal sialic acid, unlike SSEA-3, which lacks this sialic acid modification. These high-resolution views of how a glycolipid interacts with an antibody may help to advance a new class of cancer-targeting immunotherapy.
- School of Science, RMIT University, Melbourne, Victoria 3083, Australia.
Organizational Affiliation: