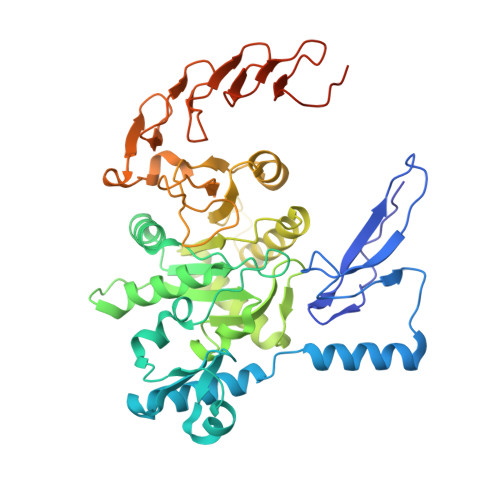
Structure and regulation of ZCCHC4 in m6A-methylation of 28S rRNA.
Ren, W., Lu, J., Huang, M., Gao, L., Li, D., Wang, G.G., Song, J.(2019) Nat Commun 10: 5042-5042
- PubMed: 31695039 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12923-x
- Primary Citation Related Structures:
6UCA - PubMed Abstract:
N 6 -methyladenosine (m 6 A) modification provides an important epitranscriptomic mechanism that critically regulates RNA metabolism and function. However, how m 6 A writers attain substrate specificities remains unclear. We report the 3.1 Å-resolution crystal structure of human CCHC zinc finger-containing protein ZCCHC4, a 28S rRNA-specific m 6 A methyltransferase, bound to S-adenosyl-L-homocysteine. The methyltransferase (MTase) domain of ZCCHC4 is packed against N-terminal GRF-type and C2H2 zinc finger domains and a C-terminal CCHC domain, creating an integrated RNA-binding surface. Strikingly, the MTase domain adopts an autoinhibitory conformation, with a self-occluded catalytic site and a fully-closed cofactor pocket. Mutational and enzymatic analyses further substantiate the molecular basis for ZCCHC4-RNA recognition and a role of the stem-loop structure within substrate in governing the substrate specificity. Overall, this study unveils unique structural and enzymatic characteristics of ZCCHC4, distinctive from what was seen with the METTL family of m 6 A writers, providing the mechanistic basis for ZCCHC4 modulation of m 6 A RNA methylation.
- Department of Biochemistry, University of California, Riverside, CA, 92521, USA.
Organizational Affiliation: