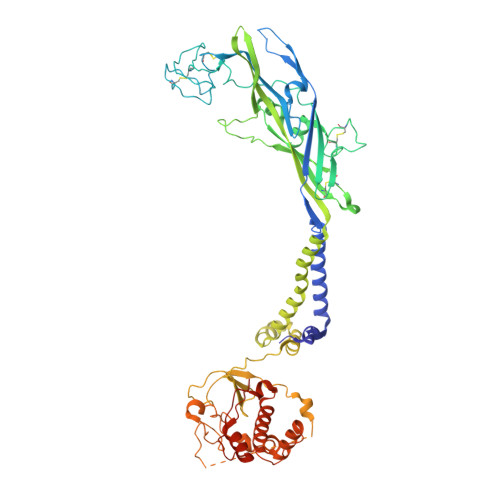
Full-Length P2X7Structures Reveal How Palmitoylation Prevents Channel Desensitization.
McCarthy, A.E., Yoshioka, C., Mansoor, S.E.(2019) Cell 179: 659-670.e13
- PubMed: 31587896 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2019.09.017
- Primary Citation Related Structures:
6U9V, 6U9W - PubMed Abstract:
P2X receptors are trimeric, non-selective cation channels activated by extracellular ATP. The P2X 7 receptor subtype is a pharmacological target because of involvement in apoptotic, inflammatory, and tumor progression pathways. It is the most structurally and functionally distinct P2X subtype, containing a unique cytoplasmic domain critical for the receptor to initiate apoptosis and not undergo desensitization. However, lack of structural information about the cytoplasmic domain has hindered understanding of the molecular mechanisms underlying these processes. We report cryoelectron microscopy structures of full-length rat P2X 7 receptor in apo and ATP-bound states. These structures reveal how one cytoplasmic element, the C-cys anchor, prevents desensitization by anchoring the pore-lining helix to the membrane with palmitoyl groups. They show a second cytoplasmic element with a unique fold, the cytoplasmic ballast, which unexpectedly contains a zinc ion complex and a guanosine nucleotide binding site. Our structures provide first insights into the architecture and function of a P2X receptor cytoplasmic domain.
- Vollum Institute, Oregon Health and Science University, Portland, OR 97239, USA; Knight Cardiovascular Institute, Oregon Health and Science University, Portland, OR 97239, USA.
Organizational Affiliation: