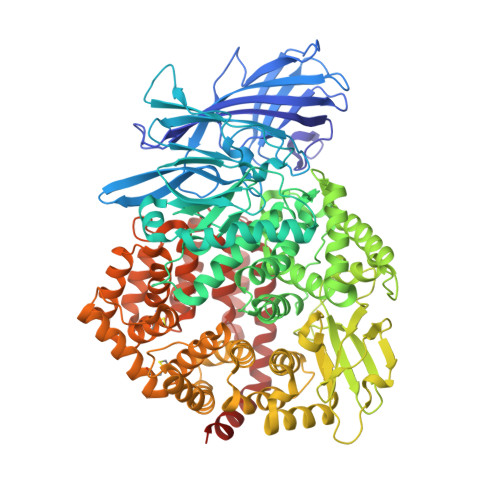
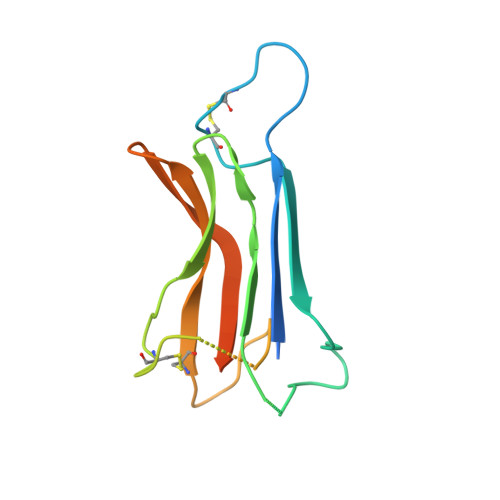
The human coronavirus HCoV-229E S-protein structure and receptor binding.
Li, Z., Tomlinson, A.C.A., Wong, A.H.M., Zhou, D., Desforges, M., Talbot, P.J., Benlekbir, S., Rubinstein, J.L., Rini, J.M.(2019) Elife 8
- PubMed: 31650956 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.51230
- Primary Citation Related Structures:
6U7E, 6U7F, 6U7G, 6U7H - PubMed Abstract:
The coronavirus S-protein mediates receptor binding and fusion of the viral and host cell membranes. In HCoV-229E, its receptor binding domain (RBD) shows extensive sequence variation but how S-protein function is maintained is not understood. Reported are the X-ray crystal structures of Class III-V RBDs in complex with human aminopeptidase N (hAPN), as well as the electron cryomicroscopy structure of the 229E S-protein. The structures show that common core interactions define the specificity for hAPN and that the peripheral RBD sequence variation is accommodated by loop plasticity. The results provide insight into immune evasion and the cross-species transmission of 229E and related coronaviruses. We also find that the 229E S-protein can expose a portion of its helical core to solvent. This is undoubtedly facilitated by hydrophilic subunit interfaces that we show are conserved among coronaviruses. These interfaces likely play a role in the S-protein conformational changes associated with membrane fusion.
- Department of Molecular Genetics, The University of Toronto, Toronto, Canada.
Organizational Affiliation: