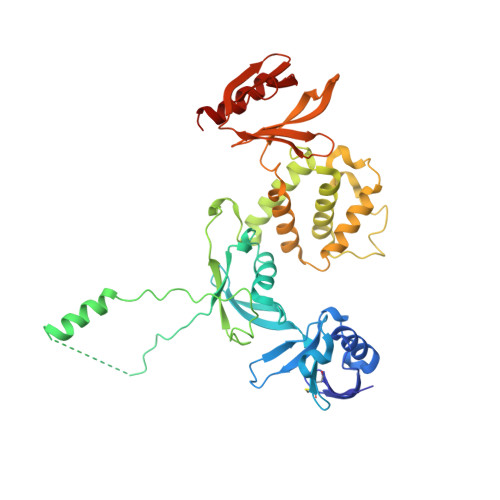
A distinct talin2 structure directs isoform specificity in cell adhesion.
Rangarajan, E.S., Primi, M.C., Colgan, L.A., Chinthalapudi, K., Yasuda, R., Izard, T.(2020) J Biological Chem 295: 12885-12899
- PubMed: 32605925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.010789
- Primary Citation Related Structures:
6U4K - PubMed Abstract:
Integrin receptors regulate normal cellular processes such as signaling, cell migration, adhesion to the extracellular matrix, and leukocyte function. Talin recruitment to the membrane is necessary for its binding to and activation of integrin. Vertebrates have two highly conserved talin homologs that differ in their expression patterns. The F1-F3 FERM subdomains of cytoskeletal proteins resemble a cloverleaf, but in talin1, its F1 subdomain and additional F0 subdomain align more linearly with its F2 and F3 subdomains. Here, we present the talin2 crystal structure, revealing that its F0-F1 di-subdomain displays another unprecedented constellation, whereby the F0-F1-F2 adopts a new cloverleaf-like arrangement. Using multiangle light scattering (MALS), fluorescence lifetime imaging (FLIM), and FRET analyses, we found that substituting the corresponding residues in talin2 that abolish lipid binding in talin1 disrupt the binding of talin to the membrane, focal adhesion formation, and cell spreading. Our results provide the molecular details of the functions of specific talin isoforms in cell adhesion.
- Cell Adhesion Laboratory, Department of Integrative Structural and Computational Biology, The Scripps Research Institute, Jupiter, Florida, USA.
Organizational Affiliation: