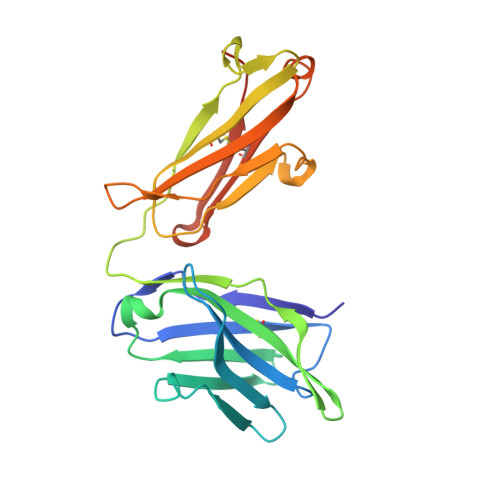
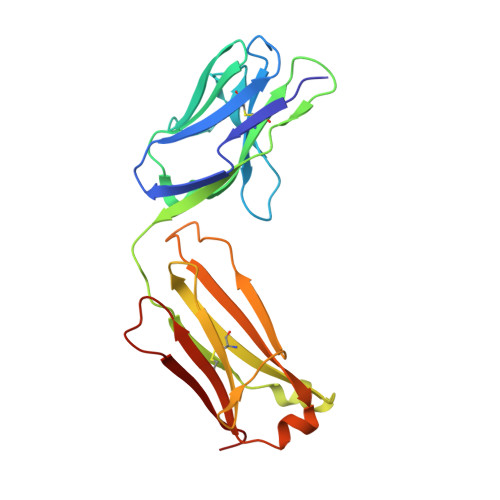
Extensive dissemination and intraclonal maturation of HIV Env vaccine-induced B cell responses.
Phad, G.E., Pushparaj, P., Tran, K., Dubrovskaya, V., Adori, M., Martinez-Murillo, P., Vazquez Bernat, N., Singh, S., Dionne, G., O'Dell, S., Bhullar, K., Narang, S., Sorini, C., Villablanca, E.J., Sundling, C., Murrell, B., Mascola, J.R., Shapiro, L., Pancera, M., Martin, M., Corcoran, M., Wyatt, R.T., Karlsson Hedestam, G.B.(2020) J Exp Medicine 217
- PubMed: 31704807
- DOI: https://doi.org/10.1084/jem.20191155
- Primary Citation Related Structures:
6U3Z, 6U9U - PubMed Abstract:
Well-ordered HIV-1 envelope glycoprotein (Env) trimers are prioritized for clinical evaluation, and there is a need for an improved understanding about how elicited B cell responses evolve following immunization. To accomplish this, we prime-boosted rhesus macaques with clade C NFL trimers and identified 180 unique Ab lineages from ∼1,000 single-sorted Env-specific memory B cells. We traced all lineages in high-throughput heavy chain (HC) repertoire (Rep-seq) data generated from multiple immune compartments and time points and expressed several as monoclonal Abs (mAbs). Our results revealed broad dissemination and high levels of somatic hypermutation (SHM) of most lineages, including tier 2 virus neutralizing lineages, following boosting. SHM was highest in the Ab complementarity determining regions (CDRs) but also surprisingly high in the framework regions (FRs), especially FR3. Our results demonstrate the capacity of the immune system to affinity-mature large numbers of Env-specific B cell lineages simultaneously, supporting the use of regimens consisting of repeated boosts to improve each Ab, even those belonging to less expanded lineages.
- Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, Stockholm, Sweden.
Organizational Affiliation: