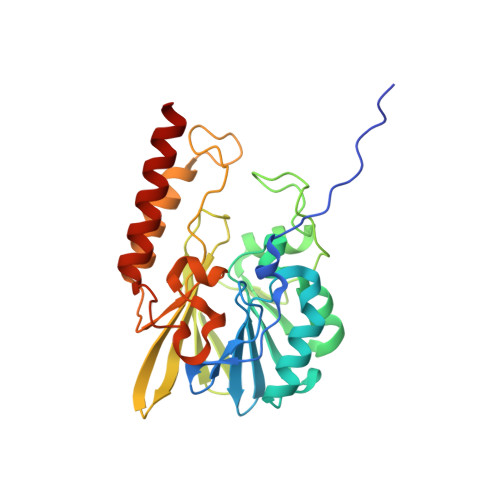
Crystal Structure of the metallo-beta-lactamase L1 from Stenotrophomonas maltophilia in the complex with the hydrolyzed moxalactam and two Ni ions
Kim, Y., Maltseva, N., Endres, M., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.