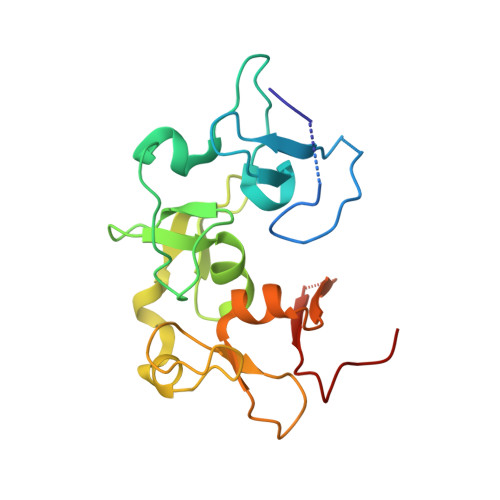
Molecular Basis for the PZP Domain of BRPF1 Association with Chromatin.
Klein, B.J., Cox, K.L., Jang, S.M., Cote, J., Poirier, M.G., Kutateladze, T.G.(2020) Structure 28: 105-110.e3
- PubMed: 31711755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.10.014
- Primary Citation Related Structures:
6U04 - PubMed Abstract:
The assembly of human histone acetyltransferase MOZ/MORF complexes relies on the scaffolding bromodomain plant homeodomain (PHD) finger 1 (BRPF1) subunit. The PHD-zinc-knuckle-PHD module of BRPF1 (BRPF1 PZP ) has been shown to associate with the histone H3 tail and DNA; however, the molecular mechanism underlying recognition of H3 and the relationship between the histone and DNA-binding activities remain unclear. In this study, we report the crystal structure of BRPF1 PZP bound to the H3 tail and characterize the role of the bipartite interaction in the engagement of BRPF1 PZP with the nucleosome core particle (NCP). We find that although both interactions of BRPF1 PZP with the H3 tail and DNA are required for tight binding to NCP and for acetyltransferase function of the BRPF1-MORF-ING5-MEAF6 complex, binding to extranucleosomal DNA dominates. Our findings suggest that functionally active BRPF1 PZP might be important in stabilization of the MOZ/MORF complexes at chromatin with accessible DNA.
- Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045, USA.
Organizational Affiliation: