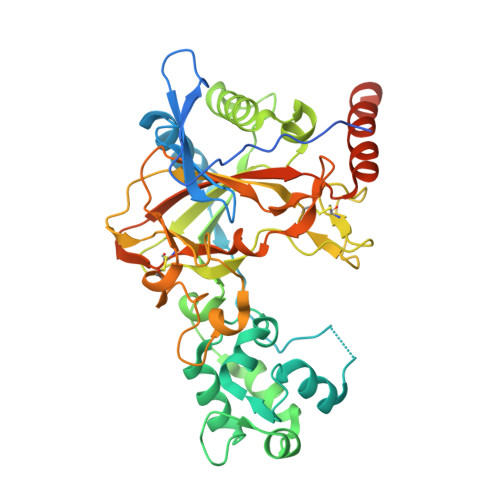
Structure of transmembrane prolyl 4-hydroxylase reveals unique organization of EF and dioxygenase domains.
Myllykoski, M., Sutinen, A., Koski, M.K., Kallio, J.P., Raasakka, A., Myllyharju, J., Wierenga, R.K., Koivunen, P.(2020) J Biological Chem 296: 100197-100197
- PubMed: 33334883 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.016542
- Primary Citation Related Structures:
6TP5 - PubMed Abstract:
Prolyl 4-hydroxylases (P4Hs) catalyze post-translational hydroxylation of peptidyl proline residues. In addition to collagen P4Hs and hypoxia-inducible factor P4Hs, a third P4H-the poorly characterized endoplasmic reticulum-localized transmembrane prolyl 4-hydroxylase (P4H-TM)-is found in animals. P4H-TM variants are associated with the familiar neurological HIDEA syndrome, but how these variants might contribute to disease is unknown. Here, we explored this question in a structural and functional analysis of soluble human P4H-TM. The crystal structure revealed an EF domain with two Ca 2+ -binding motifs inserted within the catalytic domain. A substrate-binding groove was formed between the EF domain and the conserved core of the catalytic domain. The proximity of the EF domain to the active site suggests that Ca 2+ binding is relevant to the catalytic activity. Functional analysis demonstrated that Ca 2+ -binding affinity of P4H-TM is within the range of physiological Ca 2+ concentration in the endoplasmic reticulum. P4H-TM was found both as a monomer and a dimer in the solution, but the monomer-dimer equilibrium was not regulated by Ca 2+ . The catalytic site contained bound Fe 2+ and N-oxalylglycine, which is an analogue of the cosubstrate 2-oxoglutarate. Comparison with homologous P4H structures complexed with peptide substrates showed that the substrate-interacting residues and the lid structure that folds over the substrate are conserved in P4H-TM, whereas the extensive loop structures that surround the substrate-binding groove, generating a negative surface potential, are different. Analysis of the structure suggests that the HIDEA variants cause loss of P4H-TM function. In conclusion, P4H-TM shares key structural elements with other P4Hs while having a unique EF domain.
- Biocenter Oulu, Faculty of Biochemistry and Molecular Medicine, University of Oulu, Oulu, Finland; Oulu Center for Cell-Matrix Research, University of Oulu, Oulu, Finland.
Organizational Affiliation: