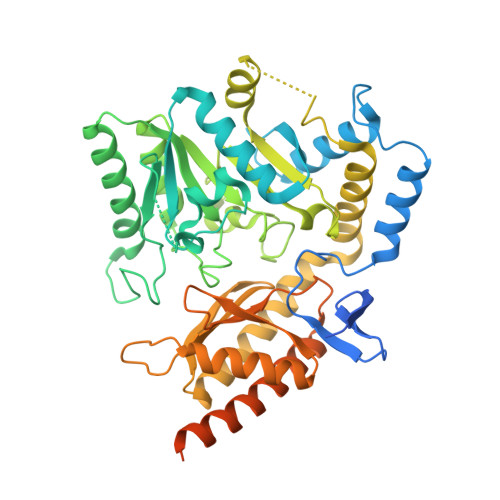
Structural characterization of human O-phosphoethanolamine phospho-lyase.
Vettraino, C., Peracchi, A., Donini, S., Parisini, E.(2020) Acta Crystallogr F Struct Biol Commun 76: 160-167
- PubMed: 32254049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20002988
- Primary Citation Related Structures:
6TOR - PubMed Abstract:
Human O-phosphoethanolamine phospho-lyase (hEtnppl; EC 4.2.3.2) is a pyridoxal 5'-phosphate-dependent enzyme that catalyzes the degradation of O-phosphoethanolamine (PEA) into acetaldehyde, phosphate and ammonia. Physiologically, the enzyme is involved in phospholipid metabolism, as PEA is the precursor of phosphatidylethanolamine in the CDP-ethanolamine (Kennedy) pathway. Here, the crystal structure of hEtnppl in complex with pyridoxamine 5'-phosphate was determined at 2.05 Å resolution by molecular replacement using the structure of A1RDF1 from Arthrobacter aurescens TC1 (PDB entry 5g4i) as the search model. Structural analysis reveals that the two proteins share the same general fold and a similar arrangement of active-site residues. These results provide novel and useful information for the complete characterization of the human enzyme.
- Center for Nano Science and Technology @PoliMi, Istituto Italiano di Tecnologia, Via Pascoli 70/3, 20133 Milano, Italy.
Organizational Affiliation: