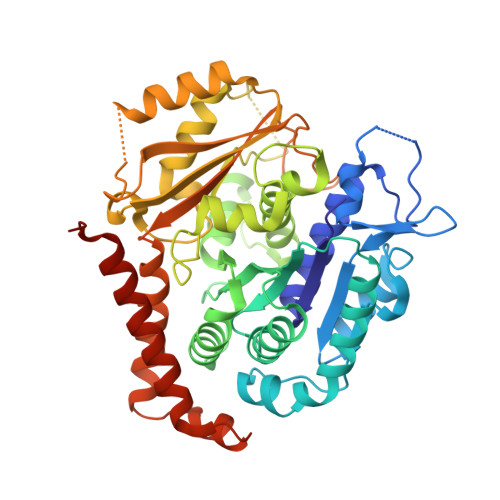
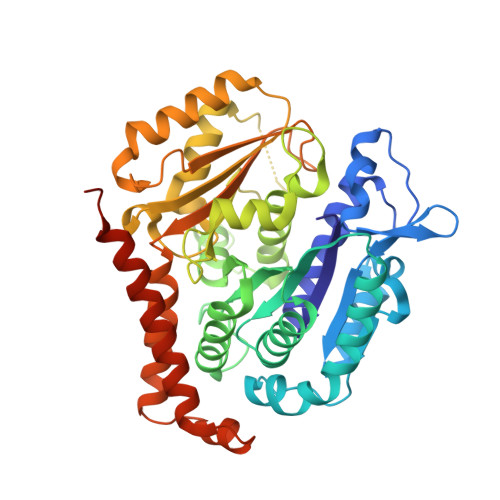
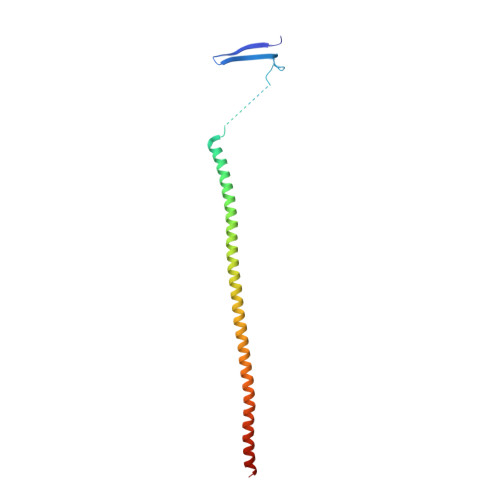
B-nor-methylene Colchicinoid PT-100 Selectively Induces Apoptosis in Multidrug-Resistant Human Cancer Cells via an Intrinsic Pathway in a Caspase-Independent Manner
Stein, A., Hilken nee Thomopoulou, P., Frias, C., Hopff, S.M., Varela, P., Wilke, N., Mariappan, A., Neudorfl, J.M., Fedorov, A.Y., Gopalakrishnan, J., Gigant, B., Prokop, A., Schmalz, H.G.(2022) ACS Omega 7: 2591-2603