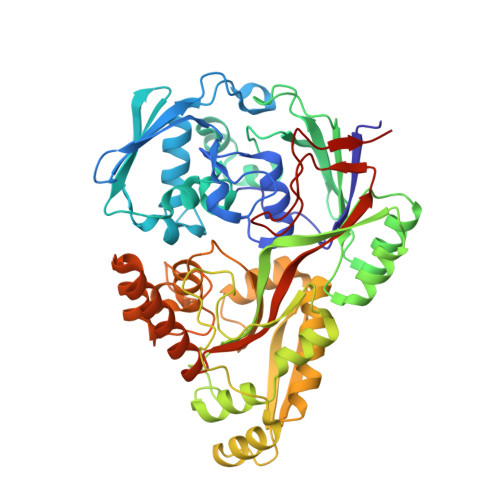
Import pathways of the mannityl-opines into the bacterial pathogen Agrobacterium tumefaciens: structural, affinity and in vivo approaches.
Vigouroux, A., Dore, J., Marty, L., Aumont-Nicaise, M., Legrand, P., Dessaux, Y., Vial, L., Morera, S.(2020) Biochem J 477: 615-628
- PubMed: 31922182 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20190886
- Primary Citation Related Structures:
6TFQ, 6TFS, 6TFX, 6TG2, 6TG3 - PubMed Abstract:
Agrobacterium tumefaciens pathogens use specific compounds denoted opines as nutrients in their plant tumor niche. These opines are produced by the host plant cells genetically modified by agrobacteria. They are imported into bacteria via solute-binding proteins (SBPs) in association with ATP-binding cassette transporters. The mannityl-opine family encompasses mannopine, mannopinic acid, agropine and agropinic acid. Structural and affinity data on mannopinic acid bound to SBPs are currently lacking while those of the three others mannityl opines are available. We investigated the molecular basis of two pathways for mannopinic acid uptake. MoaA was proposed as the specific SBP for mannopinic acid import in mannityl opines-assimilating agrobacteria, which was validated here using genetic studies and affinity measurements. We structurally characterized the mannopinic acid-binding mode of MoaA in two crystal forms at 2.05 and 1.57 Å resolution. We demonstrated that the non-specific SBP MotA, so far characterized as mannopine and Amadori compound importer, was also able to transport mannopinic acid. The structure of MotA bound to mannopinic acid at 2.2 Å resolution defines a different mannopinic acid-binding signature, similar to that of mannopine. Combining in vitro and in vivo approaches, this work allowed us to complete the characterization of the mannityl-opines assimilation pathways, highlighting the important role of two dual imports of agropinic and mannopinic acids. Our data shed new light on how the mannityl-opines contribute to the establishment of the ecological niche of agrobacteria from the early to the late stages of tumor development.
- Université Paris-Saclay, CEA, CNRS, Institute for Integrative Biology of the Cell (I2BC), 91198 Gif-sur-Yvette, France.
Organizational Affiliation: