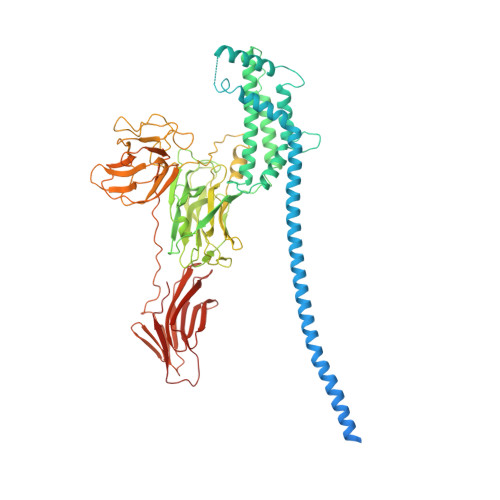
Molecular architecture and activation of the insecticidal protein Vip3Aa from Bacillus thuringiensis.
Nunez-Ramirez, R., Huesa, J., Bel, Y., Ferre, J., Casino, P., Arias-Palomo, E.(2020) Nat Commun 11: 3974-3974
- PubMed: 32769995 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-17758-5
- Primary Citation Related Structures:
6TFJ, 6TFK - PubMed Abstract:
Bacillus thuringiensis Vip3 (Vegetative Insecticidal Protein 3) toxins are widely used in biotech crops to control Lepidopteran pests. These proteins are produced as inactive protoxins that need to be activated by midgut proteases to trigger cell death. However, little is known about their three-dimensional organization and activation mechanism at the molecular level. Here, we have determined the structures of the protoxin and the protease-activated state of Vip3Aa at 2.9 Å using cryo-electron microscopy. The reconstructions show that the protoxin assembles into a pyramid-shaped tetramer with the C-terminal domains exposed to the solvent and the N-terminal region folded into a spring-loaded apex that, after protease activation, drastically remodels into an extended needle by a mechanism akin to that of influenza haemagglutinin. These results provide the molecular basis for Vip3 activation and function, and serves as a strong foundation for the development of more efficient insecticidal proteins.
- Centro de Investigaciones Biológicas Margarita Salas, CSIC, 28040, Madrid, Spain.
Organizational Affiliation: