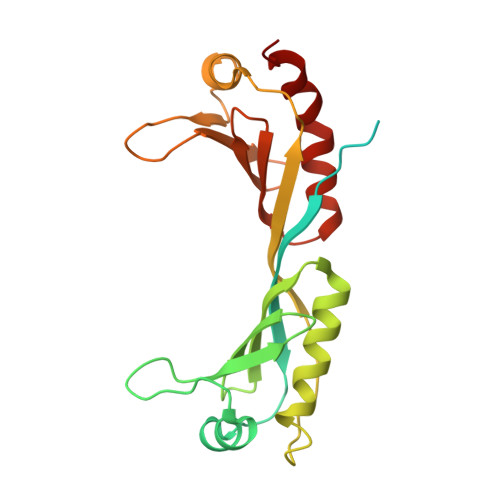
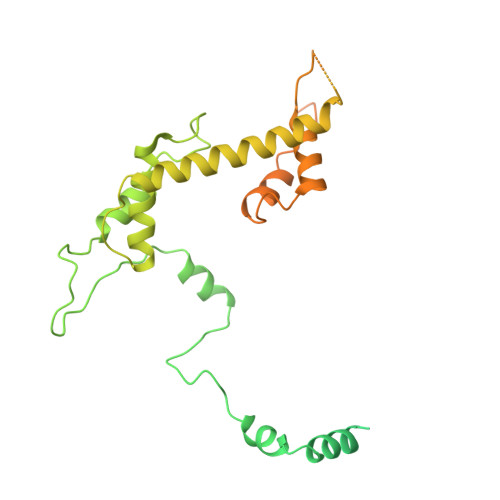
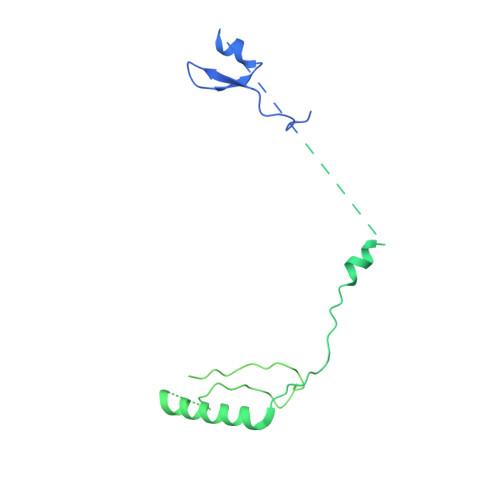
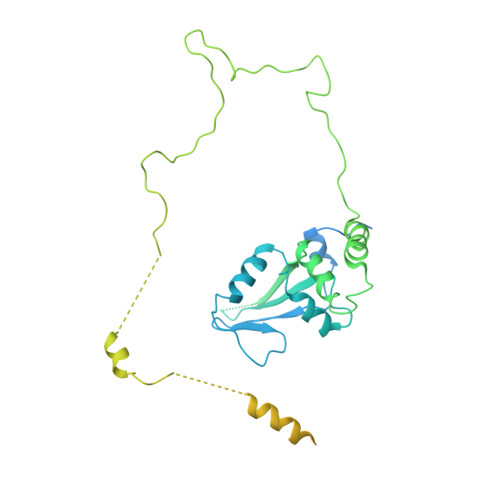
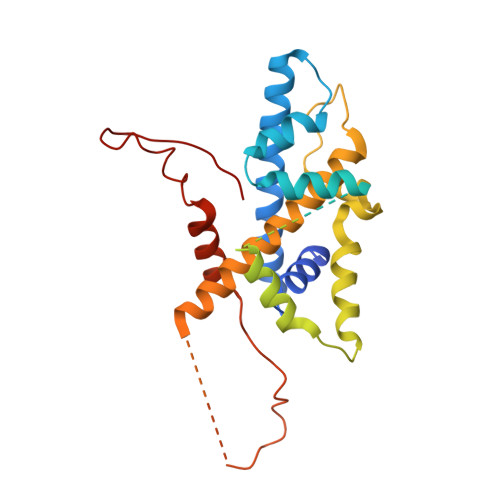
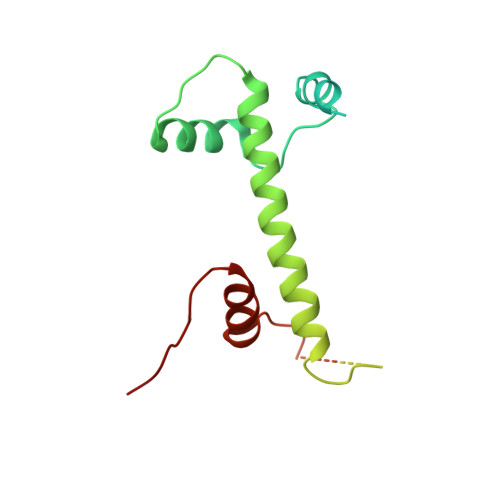
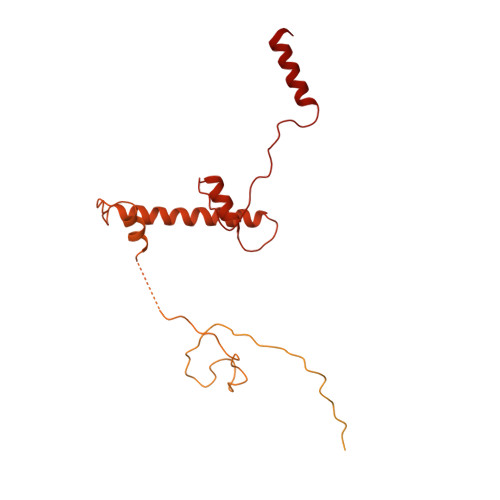
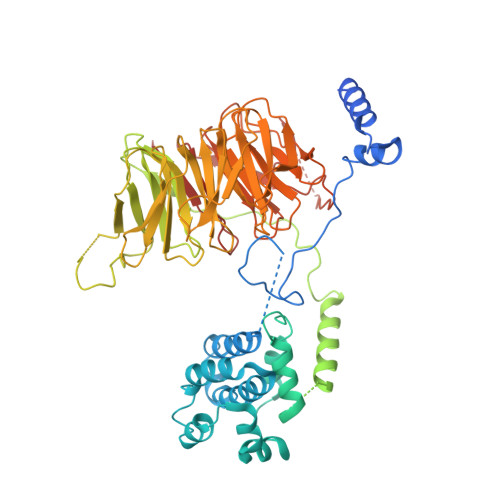
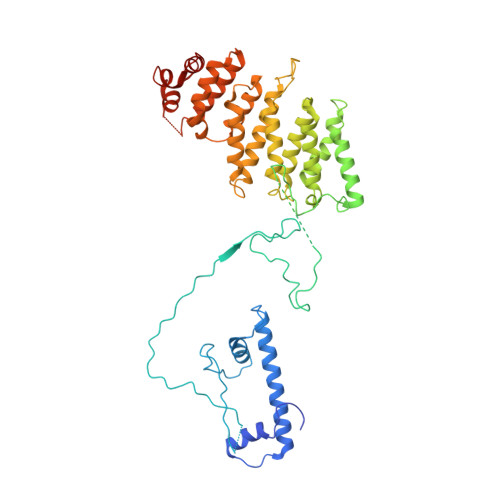
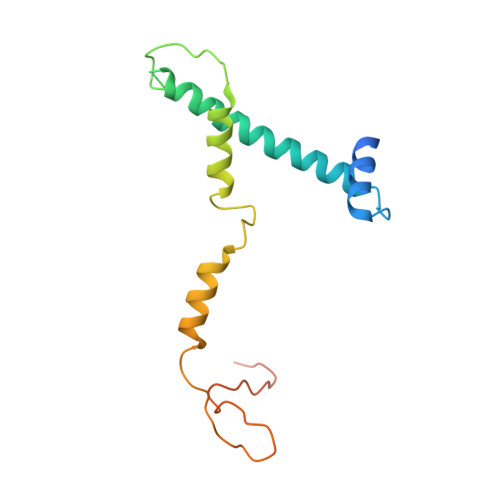
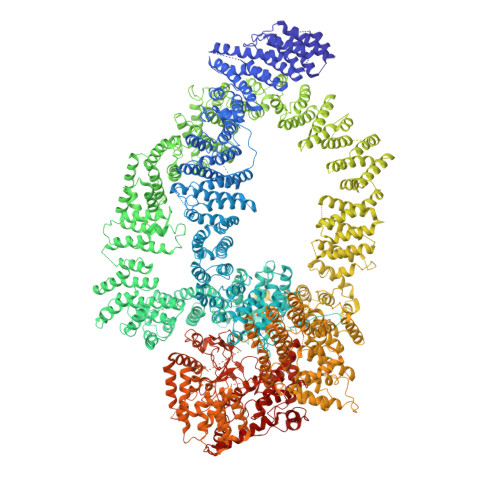
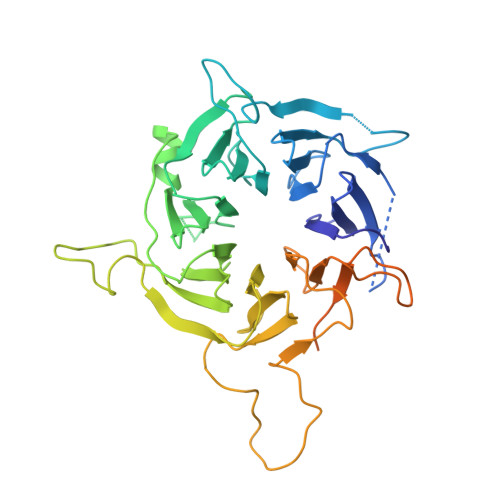
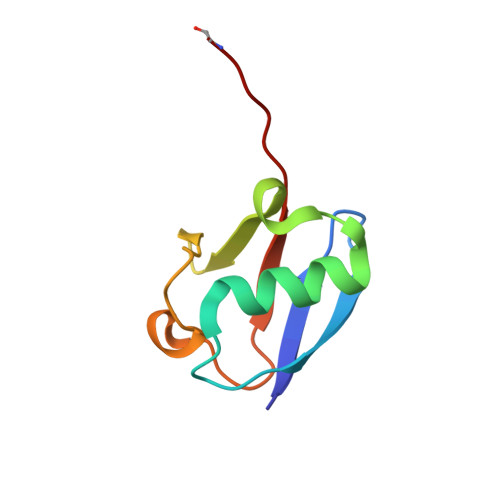
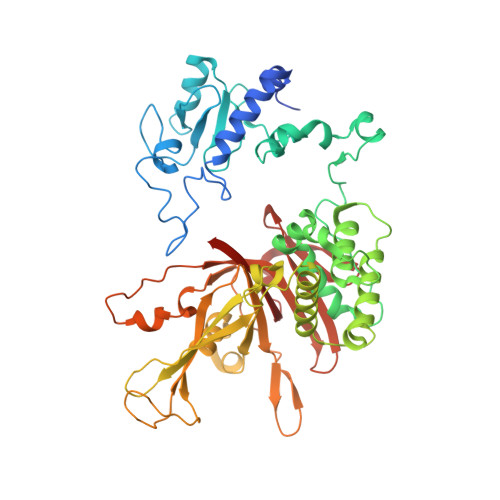
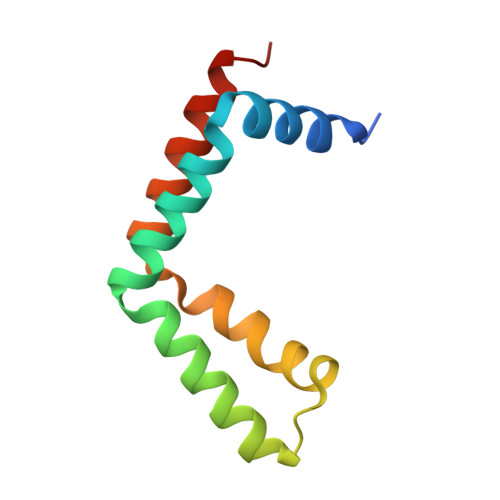
Structure of SAGA and mechanism of TBP deposition on gene promoters.
Papai, G., Frechard, A., Kolesnikova, O., Crucifix, C., Schultz, P., Ben-Shem, A.(2020) Nature 577: 711-716
- PubMed: 31969704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-1944-2
- Primary Citation Related Structures:
6TB4, 6TBM - PubMed Abstract:
SAGA (Spt-Ada-Gcn5-acetyltransferase) is a 19-subunit complex that stimulates transcription via two chromatin-modifying enzymatic modules and by delivering the TATA box binding protein (TBP) to nucleate the pre-initiation complex on DNA, a pivotal event in the expression of protein-encoding genes 1 . Here we present the structure of yeast SAGA with bound TBP. The core of the complex is resolved at 3.5 Å resolution (0.143 Fourier shell correlation). The structure reveals the intricate network of interactions that coordinate the different functional domains of SAGA and resolves an octamer of histone-fold domains at the core of SAGA. This deformed octamer deviates considerably from the symmetrical analogue in the nucleosome and is precisely tuned to establish a peripheral site for TBP, where steric hindrance represses binding of spurious DNA. Complementary biochemical analysis points to a mechanism for TBP delivery and release from SAGA that requires transcription factor IIA and whose efficiency correlates with the affinity of DNA to TBP. We provide the foundations for understanding the specific delivery of TBP to gene promoters and the multiple roles of SAGA in regulating gene expression.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire, Integrated Structural Biology Department, Equipe labellisée Ligue Contre le Cancer, Illkirch, France.
Organizational Affiliation: