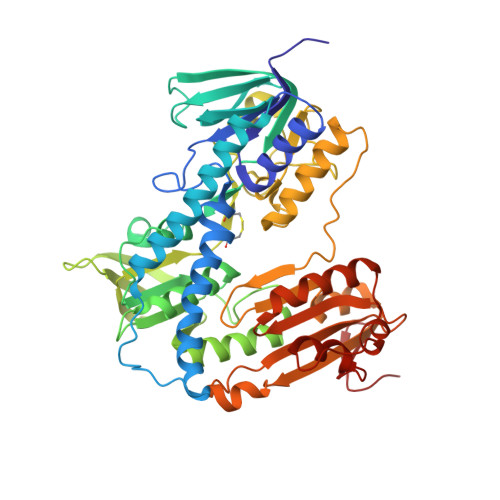
Scaffold hopping identifies new triazole-and triazolium-based inhibitors of Leishmania infantum Trypanothione Reductase with potent and selective antileishmanial activity
Revuelto, A., de Lucio, H., Carriles, A., Garcia Soriano, J.C., Sanchez-Murcia, P.A., Hermoso, J.A., Gago, F., Camarasa, M.J., Jimenez-Ruiz, A., Velazquez, S.To be published.