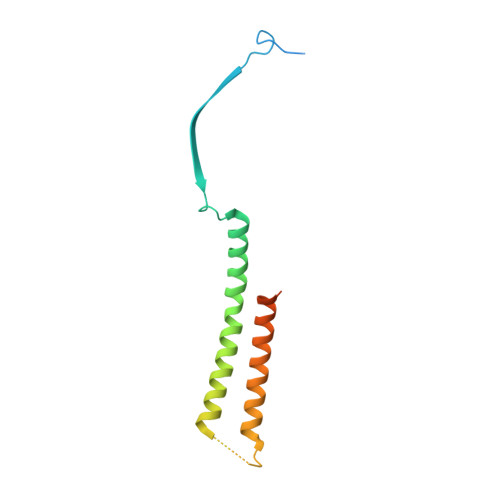
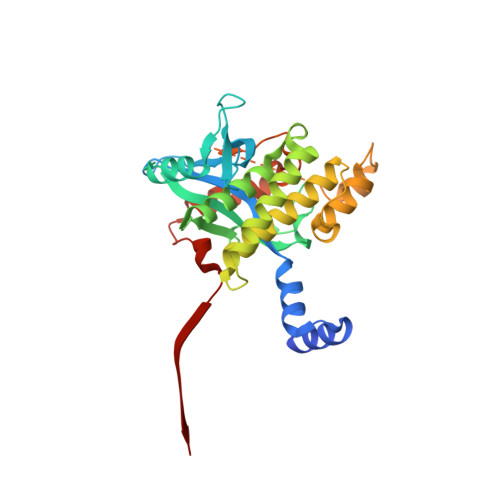
Insights into herpesvirus assembly from the structure of the pUL7:pUL51 complex.
Butt, B.G., Owen, D.J., Jeffries, C.M., Ivanova, L., Hill, C.H., Houghton, J.W., Ahmed, M.F., Antrobus, R., Svergun, D.I., Welch, J.J., Crump, C.M., Graham, S.C.(2020) Elife 9
- PubMed: 32391791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.53789
- Primary Citation Related Structures:
6T5A - PubMed Abstract:
Herpesviruses acquire their membrane envelopes in the cytoplasm of infected cells via a molecular mechanism that remains unclear. Herpes simplex virus (HSV)-1 proteins pUL7 and pUL51 form a complex required for efficient virus envelopment. We show that interaction between homologues of pUL7 and pUL51 is conserved across human herpesviruses, as is their association with trans -Golgi membranes. We characterized the HSV-1 pUL7:pUL51 complex by solution scattering and chemical crosslinking, revealing a 1:2 complex that can form higher-order oligomers in solution, and we solved the crystal structure of the core pUL7:pUL51 heterodimer. While pUL7 adopts a previously-unseen compact fold, the helix-turn-helix conformation of pUL51 resembles the cellular endosomal complex required for transport (ESCRT)-III component CHMP4B and pUL51 forms ESCRT-III-like filaments, suggesting a direct role for pUL51 in promoting membrane scission during virus assembly. Our results provide a structural framework for understanding the role of the conserved pUL7:pUL51 complex in herpesvirus assembly.
- Department of Pathology, University of Cambridge, Cambridge, United Kingdom.
Organizational Affiliation: