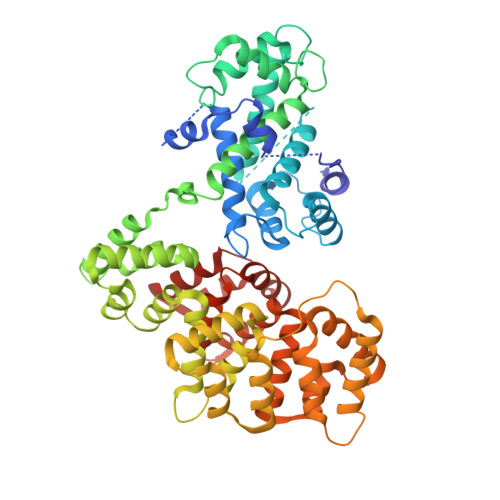
Structure Determination of the Transactivation Domain of p53 in Complex with S100A4 Using Annexin A2 as a Crystallization Chaperone.
Ecsedi, P., Gogl, G., Hof, H., Kiss, B., Harmat, V., Nyitray, L.(2020) Structure 28: 943-953.e4
- PubMed: 32442400 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2020.05.001
- Primary Citation Related Structures:
6T58 - PubMed Abstract:
To fully understand the environmental factors that influence crystallization is an enormous task, therefore crystallographers are still forced to work "blindly" trying as many crystallizing conditions and mutations to improve crystal packing as possible. Numerous times these random attempts simply fail even when using state-of-the-art techniques. As an alternative, crystallization chaperones, having good crystal-forming properties, can be invoked. Today, the almost exclusively used such protein is the maltose-binding protein (MBP) and crystallographers need other widely applicable options. Here, we introduce annexin A2 (ANXA2), which has just as good, if not better, crystal-forming ability than the wild-type MBP. Using ANXA2 as heterologous fusion partner, we were able to solve the atomic resolution structure of a challenging crystallization target, the transactivation domain (TAD) of p53 in complex with the metastasis-associated protein S100A4. p53 TAD forms an asymmetric fuzzy complex with the symmetric S1004 and could interfere with its function.
- Department of Biochemistry, ELTE Eötvös Loránd University, Budapest 1117, Hungary.
Organizational Affiliation: