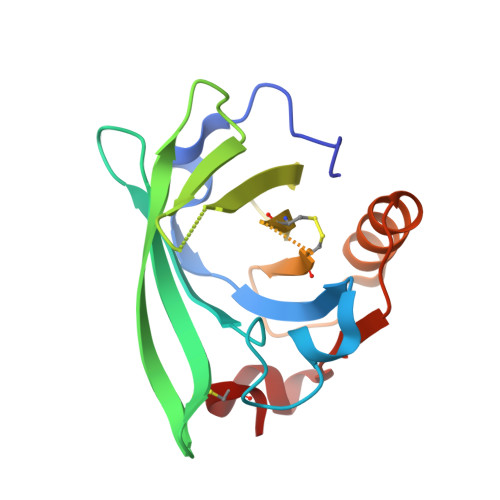
Conformational flexibility and ligand binding properties of ovine beta-lactoglobulin.
Loch, J., Bonarek, P., Lewinski, K.(2019) Acta Biochim Pol 66: 577-584
- PubMed: 31880900 Search on PubMed
- DOI: https://doi.org/10.18388/abp.2019_2883
- Primary Citation Related Structures:
6T42, 6T44 - PubMed Abstract:
Ovine β‑lactoglobulin was characterized by spectroscopic (CD), calorimetric (ITC) and X-ray structural studies. The structure of ovine β‑lactoglobulin complex with decanol showed that tight packing of molecules in the crystalline phase enforces a distortion of protein flexible loops resulting in the formation of an asymmetric dimer. The loops surrounding β-barrel in ovine lactoglobulin possessed the same conformational flexibility as observed previously in other lactoglobulins and the change of their conformation regulates the access to the binding pocket. The structure of asymmetric dimer revealed a new region in β-barrel where ligand polar group can be located. These findings indicated protein adaptability to ligand dimensions and inter- and intramolecular interactions in the crystalline phase. Calorimetric and crystallographic studies provided the experimental evidence that ovine lactoglobulin is able to bind aliphatic ligands. Thermodynamic parameters of sodium dodecyl sulfate binding determined by ITC at pH 7.5 had Ka, ΔH, TΔS and ΔG values similar to those observed for bovine and caprine protein indicating the same mechanism of ligand binding.
- Department of Crystal Chemistry and Crystal Physics, Faculty of Chemistry, Jagiellonian University, Kraków, Poland.
Organizational Affiliation: