Development of a structural epitope mimic: an idiotypic approach to HCV vaccine design.
Cowton, V.M., Owsianka, A.M., Fadda, V., Ortega-Prieto, A.M., Cole, S.J., Potter, J.A., Skelton, J.K., Jeffrey, N., Di Lorenzo, C., Dorner, M., Taylor, G.L., Patel, A.H.(2021) NPJ Vaccines 6: 7-7
- PubMed: 33420102 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41541-020-00269-1
- Primary Citation Related Structures:
6SXI - PubMed Abstract:
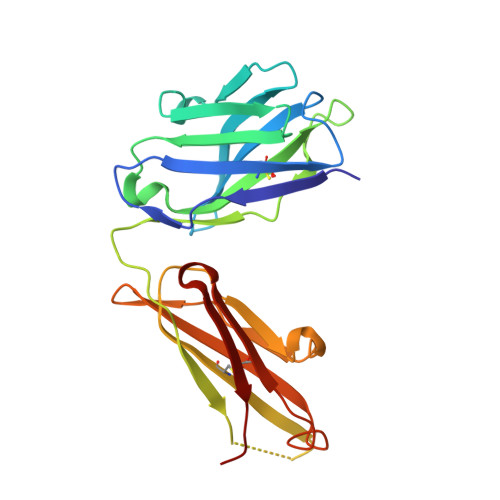
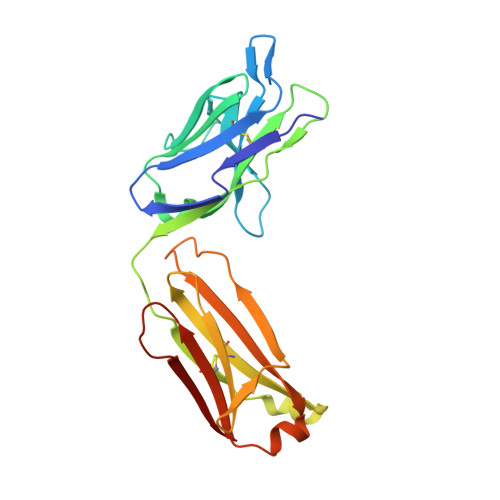
HCV vaccine development is stymied by the high genetic diversity of the virus and the variability of the envelope glycoproteins. One strategy to overcome this is to identify conserved, functionally important regions-such as the epitopes of broadly neutralizing antibodies (bNAbs)-and use these as a basis for structure-based vaccine design. Here, we report an anti-idiotype approach that has generated an antibody that mimics a highly conserved neutralizing epitope on HCV E2. Crucially, a mutagenesis screen was used to identify the antibody, designated B2.1 A, whose binding characteristics to the bNAb AP33 closely resemble those of the original antigen. Protein crystallography confirmed that B2.1 A is a structural mimic of the AP33 epitope. When used as an immunogen B2.1 A induced antibodies that recognized the same epitope and E2 residues as AP33 and most importantly protected against HCV challenge in a mouse model.
- MRC-University of Glasgow Centre for Virus Research, Garscube Campus, 464 Bearsden Road, Glasgow, UK.
Organizational Affiliation: