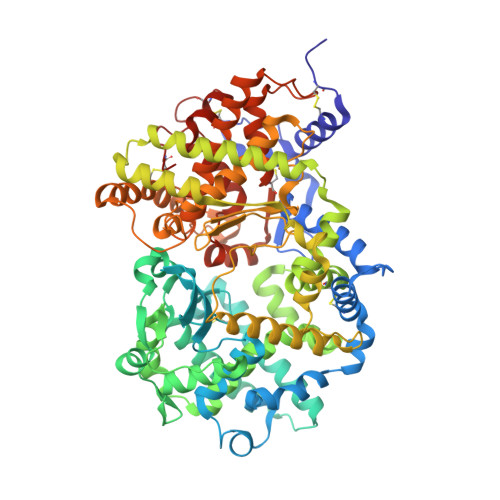
Molecular Basis for Omapatrilat and Sampatrilat Binding to Neprilysin-Implications for Dual Inhibitor Design with Angiotensin-Converting Enzyme.
Sharma, U., Cozier, G.E., Sturrock, E.D., Acharya, K.R.(2020) J Med Chem 63: 5488-5500
- PubMed: 32337993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.0c00441
- Primary Citation Related Structures:
6SUK, 6SVY, 6XVP - PubMed Abstract:
Neprilysin (NEP) and angiotensin-converting enzyme (ACE) are two key zinc-dependent metallopeptidases in the natriuretic peptide and kinin systems and renin-angiotensin-aldosterone system, respectively. They play an important role in blood pressure regulation and reducing the risk of heart failure. Vasopeptidase inhibitors omapatrilat and sampatrilat possess dual activity against these enzymes by blocking the ACE-dependent conversion of angiotensin I to the potent vasoconstrictor angiotensin II while simultaneously halting the NEP-dependent degradation of vasodilator atrial natriuretic peptide. Here, we report crystal structures of omapatrilat, sampatrilat, and sampatrilat-ASP (a sampatrilat analogue) in complex with NEP at 1.75, 2.65, and 2.6 Å, respectively. A detailed analysis of these structures and the corresponding structures of ACE with these inhibitors has provided the molecular basis of dual inhibitor recognition involving the catalytic site in both enzymes. This new information will be very useful in the design of safer and more selective vasopeptidase inhibitors of NEP and ACE for effective treatment in hypertension and heart failure.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: