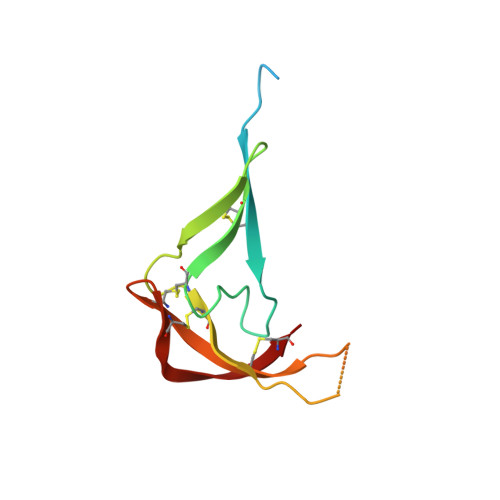
Structural characterization of anti-CCL5 activity of the tick salivary protein evasin-4.
Denisov, S.S., Ramirez-Escudero, M., Heinzmann, A.C.A., Ippel, J.H., Dawson, P.E., Koenen, R.R., Hackeng, T.M., Janssen, B.J.C., Dijkgraaf, I.(2020) J Biological Chem 295: 14367-14378
- PubMed: 32817341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.013891
- Primary Citation Related Structures:
6ST4, 6STC, 6STE, 6STK - PubMed Abstract:
Ticks, as blood-sucking parasites, have developed a complex strategy to evade and suppress host immune responses during feeding. The crucial part of this strategy is expression of a broad family of salivary proteins, called Evasins, to neutralize chemokines responsible for cell trafficking and recruitment. However, structural information about Evasins is still scarce, and little is known about the structural determinants of their binding mechanism to chemokines. Here, we studied the structurally uncharacterized Evasin-4, which neutralizes a broad range of CC-motif chemokines, including the chemokine CC-motif ligand 5 (CCL5) involved in atherogenesis. Crystal structures of Evasin-4 and E66S CCL5, an obligatory dimeric variant of CCL5, were determined to a resolution of 1.3-1.8 Å. The Evasin-4 crystal structure revealed an L-shaped architecture formed by an N- and C-terminal subdomain consisting of eight β-strands and an α-helix that adopts a substantially different position compared with closely related Evasin-1. Further investigation into E66S CCL5-Evasin-4 complex formation with NMR spectroscopy showed that residues of the N terminus are involved in binding to CCL5. The peptide derived from the N-terminal region of Evasin-4 possessed nanomolar affinity to CCL5 and inhibited CCL5 activity in monocyte migration assays. This suggests that Evasin-4 derivatives could be used as a starting point for the development of anti-inflammatory drugs.
- Department of Biochemistry, Cardiovascular Research Institute Maastricht, Maastricht University, Maastricht, The Netherlands.
Organizational Affiliation: