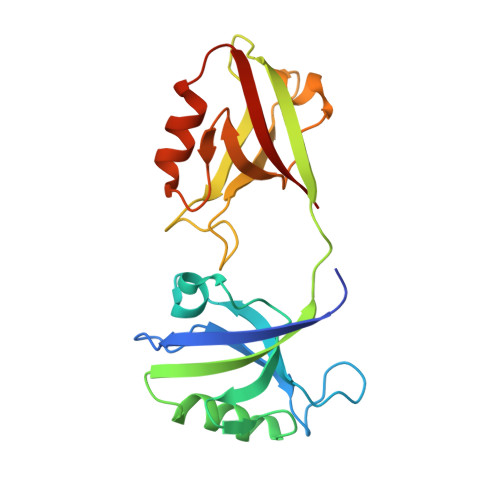
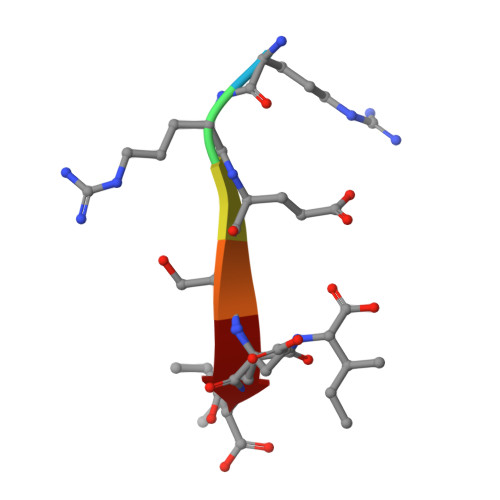
The Dual PDZ Domain from Postsynaptic Density Protein 95 Forms a Scaffold with Peptide Ligand.
Rodzli, N.A., Lockhart-Cairns, M.P., Levy, C.W., Chipperfield, J., Bird, L., Baldock, C., Prince, S.M.(2020) Biophys J 119: 667-689
- PubMed: 32652058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2020.06.018
- Primary Citation Related Structures:
6SPV, 6SPZ - PubMed Abstract:
PSD-95 is a member of the membrane-associated guanylate kinase class of proteins that forms scaffolding interactions with partner proteins, including ion and receptor channels. PSD-95 is directly implicated in modulating the electrical responses of excitable cells. The first two PSD-95/disks large/zona occludens (PDZ) domains of PSD-95 have been shown to be the key component in the formation of channel clusters. We report crystal structures of this dual domain in both apo- and ligand-bound form: thermodynamic analysis of the ligand association and small-angle x-ray scattering of the dual domain in the absence and presence of ligands. These experiments reveal that the ligated double domain forms a three-dimensional scaffold that can be described by a space group. The concentration of the components in this study is comparable with those found in compartments of excitable cells such as the postsynaptic density and juxtaparanodes of Ranvier. These in vitro experiments inform the basis of the scaffolding function of PSD-95 and provide a detailed model for scaffold formation by the PDZ domains of PSD-95.
- School of Biological Sciences, Faculty of Biology, Medicine and Health, University of Manchester, Manchester Academic Health Science Centre, Manchester, United Kingdom.
Organizational Affiliation: