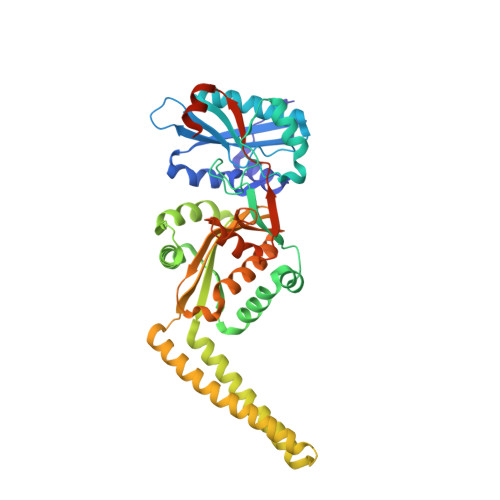
Structure-based modeling and dynamics of MurM, a Streptococcus pneumoniae penicillin resistance determinant present at the cytoplasmic membrane.
York, A., Lloyd, A.J., Del Genio, C.I., Shearer, J., Hinxman, K.J., Fritz, K., Fulop, V., Dowson, C.G., Khalid, S., Roper, D.I.(2021) Structure 29: 731-742.e6
- PubMed: 33740396 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2021.03.001
- Primary Citation Related Structures:
6SNR - PubMed Abstract:
Branched Lipid II, required for the formation of indirectly crosslinked peptidoglycan, is generated by MurM, a protein essential for high-level penicillin resistance in the human pathogen Streptococcus pneumoniae. We have solved the X-ray crystal structure of Staphylococcus aureus FemX, an isofunctional homolog, and have used this as a template to generate a MurM homology model. Using this model, we perform molecular docking and molecular dynamics to examine the interaction of MurM with the phospholipid bilayer and the membrane-embedded Lipid II substrate. Our model suggests that MurM is associated with the major membrane phospholipid cardiolipin, and experimental evidence confirms that the activity of MurM is enhanced by this phospholipid and inhibited by its direct precursor phosphatidylglycerol. The spatial association of pneumococcal membrane phospholipids and their impact on MurM activity may therefore be critical to the final architecture of peptidoglycan and the expression of clinically relevant penicillin resistance in this pathogen.
- School of Life Science, University of Warwick, Coventry, West Midlands CV4 7AL, UK.
Organizational Affiliation: