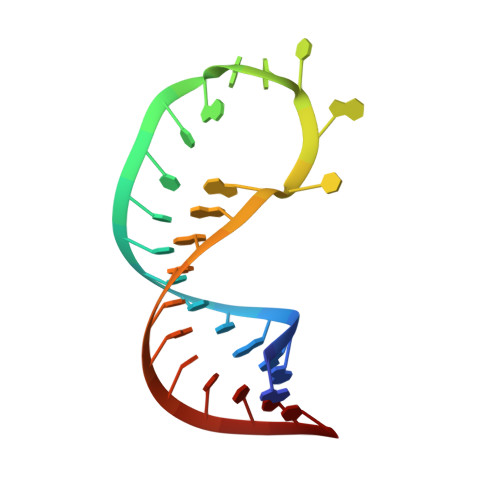
Aberrant interaction of FUS with the U1 snRNA provides a molecular mechanism of FUS induced amyotrophic lateral sclerosis.
Jutzi, D., Campagne, S., Schmidt, R., Reber, S., Mechtersheimer, J., Gypas, F., Schweingruber, C., Colombo, M., von Schroetter, C., Loughlin, F.E., Devoy, A., Hedlund, E., Zavolan, M., Allain, F.H., Ruepp, M.D.(2020) Nat Commun 11: 6341-6341
- PubMed: 33311468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-20191-3
- Primary Citation Related Structures:
6SNJ - PubMed Abstract:
Mutations in the RNA-binding protein Fused in Sarcoma (FUS) cause early-onset amyotrophic lateral sclerosis (ALS). However, a detailed understanding of central RNA targets of FUS and their implications for disease remain elusive. Here, we use a unique blend of crosslinking and immunoprecipitation (CLIP) and NMR spectroscopy to identify and characterise physiological and pathological RNA targets of FUS. We find that U1 snRNA is the primary RNA target of FUS via its interaction with stem-loop 3 and provide atomic details of this RNA-mediated mode of interaction with the U1 snRNP. Furthermore, we show that ALS-associated FUS aberrantly contacts U1 snRNA at the Sm site with its zinc finger and traps snRNP biogenesis intermediates in human and murine motor neurons. Altogether, we present molecular insights into a FUS toxic gain-of-function involving direct and aberrant RNA-binding and strengthen the link between two motor neuron diseases, ALS and spinal muscular atrophy (SMA).
- United Kingdom Dementia Research Institute Centre, Institute of Psychiatry, Psychology and Neuroscience, King's College London, Maurice Wohl Clinical Neuroscience Institute, London, UK.
Organizational Affiliation: