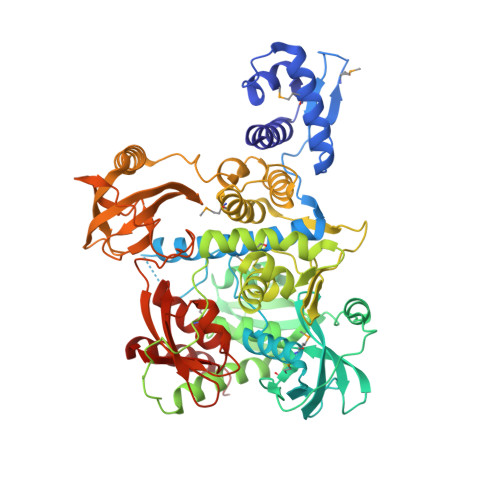
Structure and functional implications of WYL domain-containing bacterial DNA damage response regulator PafBC.
Muller, A.U., Leibundgut, M., Ban, N., Weber-Ban, E.(2019) Nat Commun 10: 4653-4653
- PubMed: 31604936 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12567-x
- Primary Citation Related Structures:
6SJ9 - PubMed Abstract:
In mycobacteria, transcriptional activator PafBC is responsible for upregulating the majority of genes induced by DNA damage. Understanding the mechanism of PafBC activation is impeded by a lack of structural information on this transcription factor that contains a widespread, but poorly understood WYL domain frequently encountered in bacterial transcription factors. Here, we determine the crystal structure of Arthrobacter aurescens PafBC. The protein consists of two modules, each harboring an N-terminal helix-turn-helix DNA-binding domain followed by a central WYL and a C-terminal extension (WCX) domain. The WYL domains exhibit Sm-folds, while the WCX domains adopt ferredoxin-like folds, both characteristic for RNA-binding proteins. Our results suggest a mechanism of regulation in which WYL domain-containing transcription factors may be activated by binding RNA or other nucleic acid molecules. Using an in vivo mutational screen in Mycobacterium smegmatis, we identify potential co-activator binding sites on PafBC.
- ETH Zurich, Institute of Molecular Biology and Biophysics, CH-8093, Zurich, Switzerland.
Organizational Affiliation: