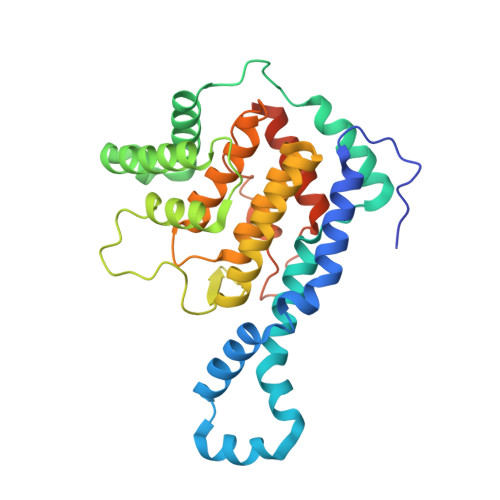
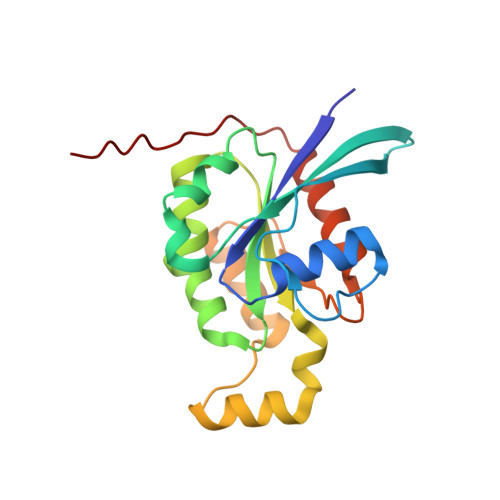
Identification of targets of AMPylating Fic enzymes by co-substrate-mediated covalent capture.
Gulen, B., Rosselin, M., Fauser, J., Albers, M.F., Pett, C., Krisp, C., Pogenberg, V., Schluter, H., Hedberg, C., Itzen, A.(2020) Nat Chem 12: 732-739
- PubMed: 32632184 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41557-020-0484-6
- Primary Citation Related Structures:
6SIU - PubMed Abstract:
Various pathogenic bacteria use post-translational modifications to manipulate the central components of host cell functions. Many of the enzymes released by these bacteria belong to the large Fic family, which modify targets with nucleotide monophosphates. The lack of a generic method for identifying the cellular targets of Fic family enzymes hinders investigation of their role and the effect of the post-translational modification. Here, we establish an approach that uses reactive co-substrate-linked enzymes for proteome profiling. We combine synthetic thiol-reactive nucleotide derivatives with recombinantly produced Fic enzymes containing strategically placed cysteines in their active sites to yield reactive binary probes for covalent substrate capture. The binary complexes capture their targets from cell lysates and permit subsequent identification. Furthermore, we determined the structures of low-affinity ternary enzyme-nucleotide-substrate complexes by applying a covalent-linking strategy. This approach thus allows target identification of the Fic enzymes from both bacteria and eukarya.
- Center for Integrated Protein Science Munich (CIPSM), Department of Chemistry, Technical University of Munich, Garching, Germany.
Organizational Affiliation: