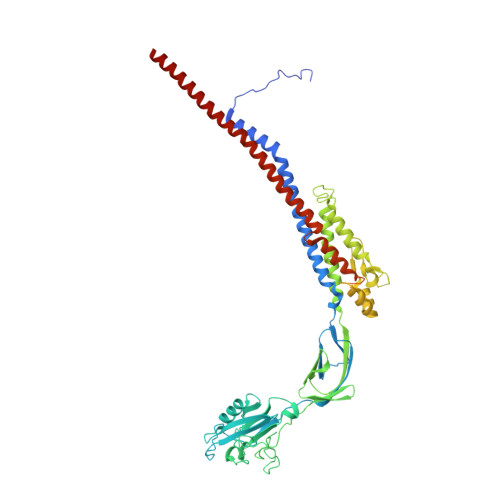
The cryo-EM structure of the bacterial flagellum cap complex suggests a molecular mechanism for filament elongation.
Al-Otaibi, N.S., Taylor, A.J., Farrell, D.P., Tzokov, S.B., DiMaio, F., Kelly, D.J., Bergeron, J.R.C.(2020) Nat Commun 11: 3210-3210
- PubMed: 32587243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-16981-4
- Primary Citation Related Structures:
6SIH - PubMed Abstract:
The bacterial flagellum is a remarkable molecular motor, whose primary function in bacteria is to facilitate motility through the rotation of a filament protruding from the bacterial cell. A cap complex, consisting of an oligomer of the protein FliD, is localized at the tip of the flagellum, and is essential for filament assembly, as well as adherence to surfaces in some bacteria. However, the structure of the intact cap complex, and the molecular basis for its interaction with the filament, remains elusive. Here we report the cryo-EM structure of the Campylobacter jejuni cap complex, which reveals that FliD is pentameric, with the N-terminal region of the protomer forming an extensive set of contacts across several subunits, that contribute to FliD oligomerization. We also demonstrate that the native C. jejuni flagellum filament is 11-stranded, contrary to a previously published cryo-EM structure, and propose a molecular model for the filament-cap interaction.
- Department of Molecular Biology and Biotechnology, the University of Sheffield, Sheffield, UK.
Organizational Affiliation: