Structure-Function Characterization of the Conserved Regulatory Mechanism of the Escherichia coli M48 Metalloprotease BepA.
Bryant, J.A., Cadby, I.T., Chong, Z.S., Boelter, G., Sevastsyanovich, Y.R., Morris, F.C., Cunningham, A.F., Kritikos, G., Meek, R.W., Banzhaf, M., Chng, S.S., Lovering, A.L., Henderson, I.R.(2020) J Bacteriol 203
- PubMed: 33106348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00434-20
- Primary Citation Related Structures:
6SAR - PubMed Abstract:
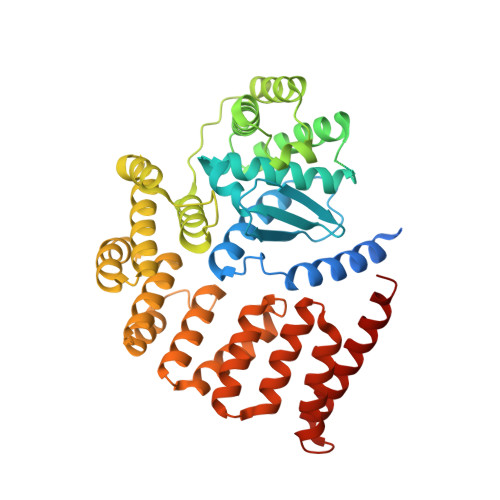
The asymmetric Gram-negative outer membrane (OM) is the first line of defense for bacteria against environmental insults and attack by antimicrobials. The key component of the OM is lipopolysaccharide, which is transported to the surface by the essential lipopolysaccharide transport (Lpt) system. Correct folding of the Lpt system component LptD is regulated by a periplasmic metalloprotease, BepA. Here, we present the crystal structure of BepA from Escherichia coli , solved to a resolution of 2.18 Å, in which the M48 protease active site is occluded by an active-site plug. Informed by our structure, we demonstrate that free movement of the active-site plug is essential for BepA function, suggesting that the protein is autoregulated by the active-site plug, which is conserved throughout the M48 metalloprotease family. Targeted mutagenesis of conserved residues reveals that the negative pocket and the tetratricopeptide repeat (TPR) cavity are required for function and degradation of the BAM complex component BamA under conditions of stress. Last, we show that loss of BepA causes disruption of OM lipid asymmetry, leading to surface exposed phospholipid. IMPORTANCE M48 metalloproteases are widely distributed in all domains of life. E. coli possesses four members of this family located in multiple cellular compartments. The functions of these proteases are not well understood. Recent investigations revealed that one family member, BepA, has an important role in the maturation of a central component of the lipopolysaccharide (LPS) biogenesis machinery. Here, we present the structure of BepA and the results of a structure-guided mutagenesis strategy, which reveal the key residues required for activity that inform how all M48 metalloproteases function.
- Institute of Microbiology and Infection, University of Birmingham, Edgbaston, United Kingdom j.a.bryant@bham.ac.uk i.henderson@imb.uq.edu.au.
Organizational Affiliation: