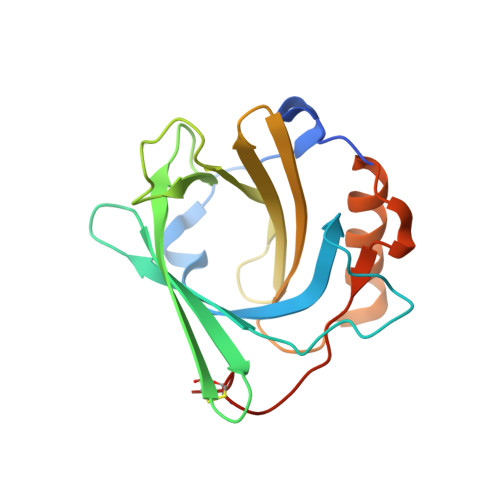
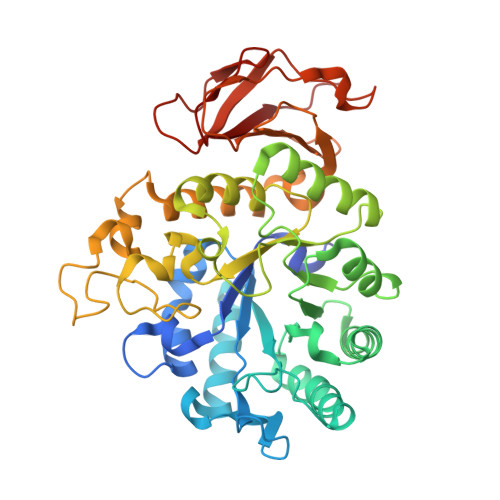
Development of a high affinity Anticalin®directed against human CD98hc for theranostic applications.
Deuschle, F.C., Morath, V., Schiefner, A., Brandt, C., Ballke, S., Reder, S., Steiger, K., Schwaiger, M., Weber, W., Skerra, A.(2020) Theranostics 10: 2172-2187
- PubMed: 32089738 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7150/thno.38968
- Primary Citation Related Structures:
6S8V - PubMed Abstract:
Enhanced amino acid supply and dysregulated integrin signaling constitute two hallmarks of cancer and are pivotal for metastatic transformation of cells. In line with its function at the crossroads of both processes, overexpression of CD98hc is clinically observed in various cancer malignancies, thus rendering it a promising tumor target. Methods : We describe the development of Anticalin proteins based on the lipocalin 2 (Lcn2) scaffold against the human CD98hc ectodomain (hCD98hcED) using directed evolution and protein design. X-ray structural analysis was performed to identify the epitope recognized by the lead Anticalin candidate. The Anticalin - with a tuned plasma half-life using PASylation ® technology - was labeled with 89 Zr and investigated by positron emission tomography (PET) of CD98-positive tumor xenograft mice. Results : The Anticalin P3D11 binds CD98hc with picomolar affinity and recognizes a protruding loop structure surrounded by several glycosylation sites within the solvent exposed membrane-distal part of the hCD98hcED. In vitro studies revealed specific binding activity of the Anticalin towards various CD98hc-expressing human tumor cell lines, suggesting broader applicability in cancer research. PET/CT imaging of mice bearing human prostate carcinoma xenografts using the optimized and 89 Zr-labeled Anticalin demonstrated strong and specific tracer accumulation (8.6 ± 1.1 %ID/g) as well as a favorable tumor-to-blood ratio of 11.8. Conclusion : Our findings provide a first proof of concept to exploit CD98hc for non-invasive biomedical imaging. The novel Anticalin-based αhCD98hc radiopharmaceutical constitutes a promising tool for preclinical and, potentially, clinical applications in oncology.
- Lehrstuhl für Biologische Chemie, Technische Universität München, 85354 Freising, Germany.
Organizational Affiliation: