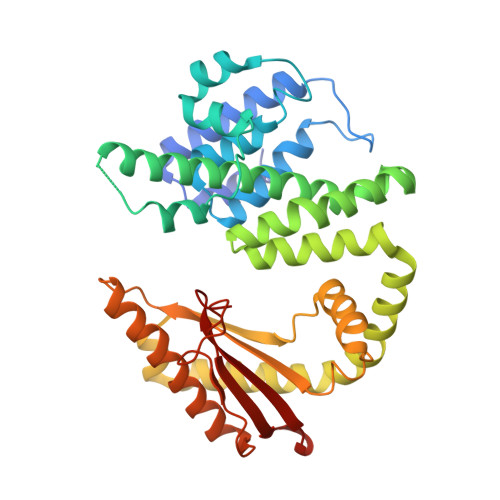
A nucleotide-switch mechanism mediates opposing catalytic activities of Rel enzymes.
Tamman, H., Van Nerom, K., Takada, H., Vandenberk, N., Scholl, D., Polikanov, Y., Hofkens, J., Talavera, A., Hauryliuk, V., Hendrix, J., Garcia-Pino, A.(2020) Nat Chem Biol 16: 834-840
- PubMed: 32393900 Search on PubMed
- DOI: https://doi.org/10.1038/s41589-020-0520-2
- Primary Citation Related Structures:
6S2T, 6S2U, 6S2V - PubMed Abstract:
Bifunctional Rel stringent factors, the most abundant class of RelA/SpoT homologs, are ribosome-associated enzymes that transfer a pyrophosphate from ATP onto the 3' of guanosine tri-/diphosphate (GTP/GDP) to synthesize the bacterial alarmone (p)ppGpp, and also catalyze the 3' pyrophosphate hydrolysis to degrade it. The regulation of the opposing activities of Rel enzymes is a complex allosteric mechanism that remains an active research topic despite decades of research. We show that a guanine-nucleotide-switch mechanism controls catalysis by Thermus thermophilus Rel (Rel Tt ). The binding of GDP/ATP opens the N-terminal catalytic domains (NTD) of Rel Tt (Rel Tt NTD ) by stretching apart the two catalytic domains. This activates the synthetase domain and allosterically blocks hydrolysis. Conversely, binding of ppGpp to the hydrolase domain closes the NTD, burying the synthetase active site and precluding the binding of synthesis precursors. This allosteric mechanism is an activity switch that safeguards against futile cycles of alarmone synthesis and degradation.
- Cellular and Molecular Microbiology, Faculté des Sciences, Université Libre de Bruxelles, Brussels, Belgium.
Organizational Affiliation: