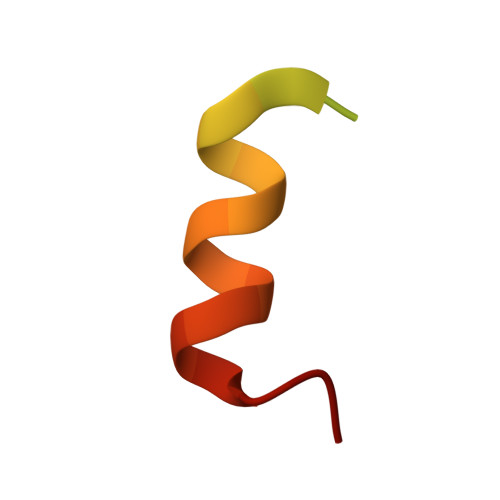
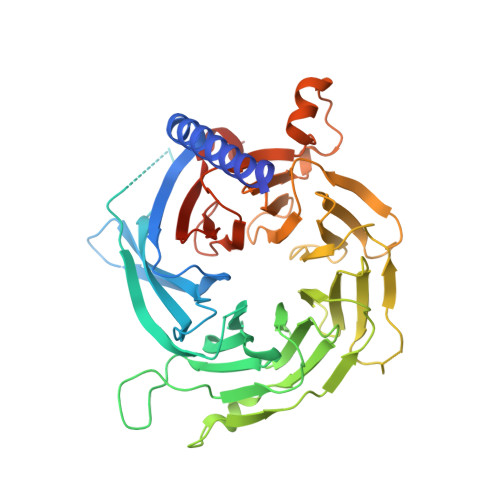
Subunit interactions and arrangements in the fission yeast Mis16-Mis18-Mis19 complex.
Korntner-Vetter, M., Lefevre, S., Hu, X.W., George, R., Singleton, M.R.(2019) Life Sci Alliance 2
- PubMed: 31371524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.201900408
- Primary Citation Related Structures:
6S1L, 6S1R, 6S29 - PubMed Abstract:
Centromeric chromatin in fission yeast is distinguished by the presence of nucleosomes containing the histone H3 variant Cnp1 CENP-A Cell cycle-specific deposition of Cnp1 requires the Mis16-Mis18-Mis19 complex, which is thought to direct recruitment of Scm3-chaperoned Cnp1/histone H4 dimers to DNA. Here, we present the structure of the essential Mis18 partner protein Mis19 and describe its interaction with Mis16, revealing a bipartite-binding site. We provide data on the stoichiometry and overall architecture of the complex and provide detailed insights into the Mis18-Mis19 interface.
- Structural Biology of Chromosome Segregation Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: