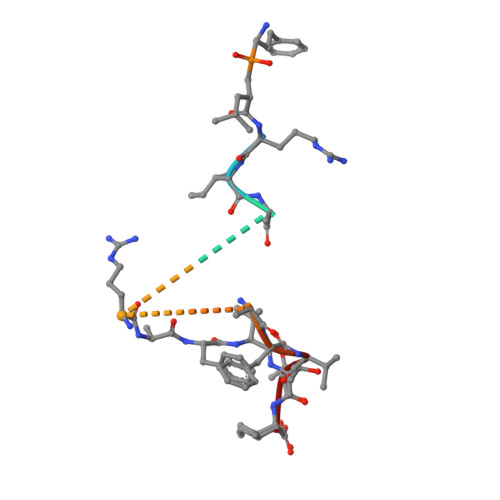
Mechanism for antigenic peptide selection by endoplasmic reticulum aminopeptidase 1.
Giastas, P., Mpakali, A., Papakyriakou, A., Lelis, A., Kokkala, P., Neu, M., Rowland, P., Liddle, J., Georgiadis, D., Stratikos, E.(2019) Proc Natl Acad Sci U S A
- PubMed: 31843903 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1912070116
- Primary Citation Related Structures:
6RQX, 6RYF - PubMed Abstract:
Endoplasmic reticulum aminopeptidase 1 (ERAP1) is an intracellular enzyme that optimizes the peptide cargo of major histocompatibility class I (MHC-I) molecules and regulates adaptive immunity. It has unusual substrate selectivity for length and sequence, resulting in poorly understood effects on the cellular immunopeptidome. To understand substrate selection by ERAP1, we solved 2 crystal structures of the enzyme with bound transition-state pseudopeptide analogs at 1.68 Å and 1.72 Å. Both peptides have their N terminus bound at the active site and extend away along a large internal cavity, interacting with shallow pockets that can influence selectivity. The longer peptide is disordered through the central region of the cavity and has its C terminus bound in an allosteric pocket of domain IV that features a carboxypeptidase-like structural motif. These structures, along with enzymatic and computational analyses, explain how ERAP1 can select peptides based on length while retaining the broad sequence-specificity necessary for its biological function.
- Institute of Nuclear & Radiological Sciences and Technology, Energy & Safety, National Centre for Scientific Research Demokritos, Agia Paraskevi, Athens 15341, Greece.
Organizational Affiliation: