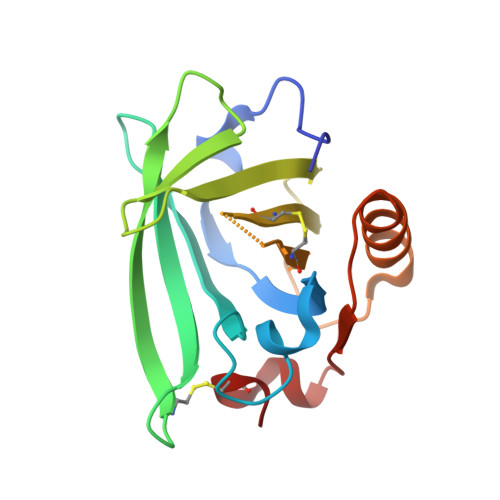
Structure-based design approach to rational site-directed mutagenesis of beta-lactoglobulin.
Bonarek, P., Loch, J.I., Tworzydlo, M., Cooper, D.R., Milto, K., Wrobel, P., Kurpiewska, K., Lewinski, K.(2020) J Struct Biol 210: 107493-107493
- PubMed: 32169624 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2020.107493
- Primary Citation Related Structures:
6QI6, 6QI7, 6QPD, 6QPE, 6RWP, 6RWQ, 6RWR, 6RYT, 6XVE - PubMed Abstract:
Recombinant proteins play an important role in medicine and have diverse applications in industrial biotechnology. Lactoglobulin has shown great potential for use in targeted drug delivery and body fluid detoxification because of its ability to bind a variety of molecules. In order to modify the biophysical properties of β-lactoglobulin, a series of single-site mutations were designed using a structure-based approach. A 3-dimensional structure alignment of homologous molecules led to the design of nine β-lactoglobulin variants with mutations introduced in the binding pocket region. Seven stable and correctly folded variants (L39Y, I56F, L58F, V92F, V92Y, F105L, M107L) were thoroughly characterized by fluorescence, circular dichroism, isothermal titration calorimetry, size-exclusion chromatography, and X-ray structural investigations. The effects of the amino acid substitutions were observed as slight rearrangements of the binding pocket geometry, but they also significantly influenced the global properties of the protein. Most of the mutations increased the thermal/chemical stability without altering the dimerization constant or pH-dependent conformational behavior. The crystal structures reveal that the I56F and F105L mutations reduced the depth of the binding pocket, which is advantageous since it can reduce the affinity to endogenous fatty acids. The F105L mutant created a unique binding mode for a fatty acid, supporting the idea that lactoglobulin can be altered to bind unique molecules. Selected variants possessing a unique combination of their individual properties can be used for further, more advanced mutagenesis, and the presented results support further research using β-lactoglobulin as a therapeutic delivery agent or a blood detoxifying molecule.
- Jagiellonian University, Faculty of Biochemistry, Biophysics and Biotechnology, Gronostajowa 7, 30-387 Kraków, Poland.
Organizational Affiliation: