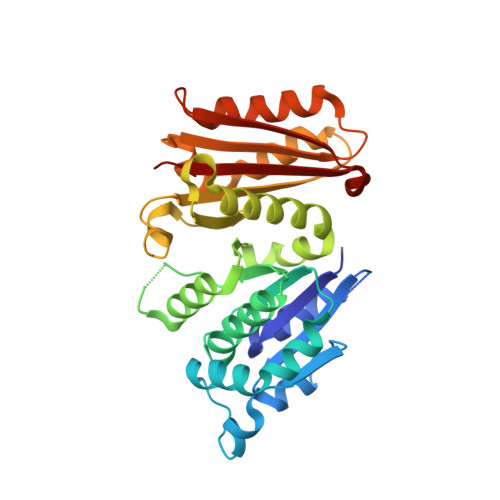
Nucleotide-induced folding of cell division protein FtsZ from Staphylococcus aureus.
Huecas, S., Canosa-Valls, A.J., Araujo-Bazan, L., Ruiz, F.M., Laurents, D.V., Fernandez-Tornero, C., Andreu, J.M.(2020) FEBS J 287: 4048-4067
- PubMed: 31997533 Search on PubMed
- DOI: https://doi.org/10.1111/febs.15235
- Primary Citation Related Structures:
6RVM, 6RVN, 6RVP, 6RVQ, 6SI9 - PubMed Abstract:
The essential bacterial division protein FtsZ uses GTP binding and hydrolysis to assemble into dynamic filaments that treadmill around the Z-ring, guiding septal wall synthesis and cell division. FtsZ is a structural homolog of tubulin and a target for discovering new antibiotics. Here, using FtsZ from the pathogen S. aureus (SaFtsZ), we reveal that, prior to assembly, FtsZ monomers require nucleotide binding for folding; this is possibly relevant to other mesophilic FtsZs. Apo-SaFtsZ is essentially unfolded, as assessed by nuclear magnetic resonance and circular dichroism. Binding of GTP (≥ 1 mm) dramatically shifts the equilibrium toward the active folded protein. Supportingly, SaFtsZ refolded with GDP crystallizes in a native structure. Apo-SaFtsZ also folds with 3.4 m glycerol, enabling high-affinity GTP binding (K D 20 nm determined by isothermal titration calorimetry) similar to thermophilic stable FtsZ. Other stabilizing agents that enhance nucleotide binding include ethylene glycol, trimethylamine N-oxide, and several bacterial osmolytes. High salt stabilizes SaFtsZ without bound nucleotide in an inactive twisted conformation. We identified a cavity behind the SaFtsZ-GDP nucleotide-binding pocket that harbors different small compounds, which is available for extended nucleotide-replacing inhibitors. Furthermore, we devised a competition assay to detect any inhibitors that overlap the nucleotide site of SaFtsZ, or Escherichia coli FtsZ, employing osmolyte-stabilized apo-FtsZs and the specific fluorescence anisotropy change in mant-GTP upon dissociation from the protein. This robust assay provides a basis to screening for high-affinity GTP-replacing ligands, which combined with structural studies and phenotypic profiling should facilitate development of a next generation of FtsZ-targeting antibacterial inhibitors.
- Centro de Investigaciones Biológicas Margarita Salas CSIC, Madrid, Spain.
Organizational Affiliation: