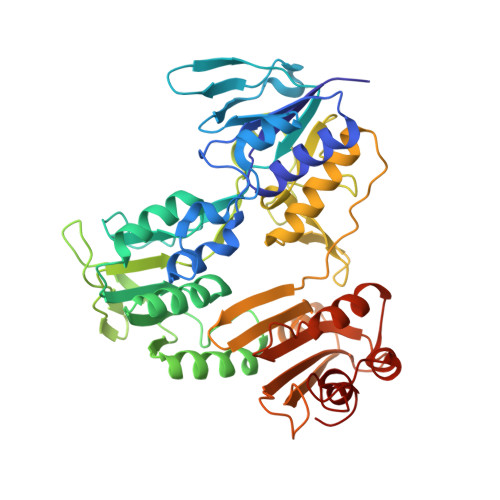
Characterization and X-ray structure of the NADH-dependent coenzyme A disulfide reductase from Thermus thermophilus.
Lencina, A.M., Koepke, J., Preu, J., Muenke, C., Gennis, R.B., Michel, H., Schurig-Briccio, L.A.(2019) Biochim Biophys Acta Bioenerg 1860: 148080-148080
- PubMed: 31520616
- DOI: https://doi.org/10.1016/j.bbabio.2019.148080
- Primary Citation Related Structures:
6RUZ, 6RVB, 6RVH - PubMed Abstract:
The crystal structure of the enzyme previously characterized as a type-2 NADH:menaquinone oxidoreductase (NDH-2) from Thermus thermophilus has been solved at a resolution of 2.9 Å and revealed that this protein is, in fact, a coenzyme A-disulfide reductase (CoADR). Coenzyme A (CoASH) replaces glutathione as the major low molecular weight thiol in Thermus thermophilus and is maintained in the reduced state by this enzyme (CoADR). Although the enzyme does exhibit NADH:menadione oxidoreductase activity expected for NDH-2 enzymes, the specific activity with CoAD as an electron acceptor is about 5-fold higher than with menadione. Furthermore, the crystal structure contains coenzyme A covalently linked Cys44, a catalytic intermediate (Cys44-S-S-CoA) reduced by NADH via the FAD cofactor. Soaking the crystals with menadione shows that menadione can bind to a site near the redox active FAD, consistent with the observed NADH:menadione oxidoreductase activity. CoADRs from other species were also examined and shown to have measurable NADH:menadione oxidoreductase activity. Although a common feature of this family of enzymes, no biological relevance is proposed. The CoADR from T. thermophilus is a soluble homodimeric enzyme. Expression of the recombinant TtCoADR at high levels in E. coli results in a small fraction that co-purifies with the membrane fraction, which was used previously to isolate the enzyme wrongly identified as a membrane-bound NDH-2. It is concluded that T. thermophilus does not contain an authentic NDH-2 component in its aerobic respiratory chain.
- Department of Biochemistry, University of Illinois, 600 S. Mathews Street, Urbana, IL 61801, USA.
Organizational Affiliation: