Insights into an unusual Auxiliary Activity 9 family member lacking the histidine brace motif of lytic polysaccharide monooxygenases.
Frandsen, K.E.H., Tovborg, M., Jorgensen, C.I., Spodsberg, N., Rosso, M.N., Hemsworth, G.R., Garman, E.F., Grime, G.W., Poulsen, J.N., Batth, T.S., Miyauchi, S., Lipzen, A., Daum, C., Grigoriev, I.V., Johansen, K.S., Henrissat, B., Berrin, J.G., Lo Leggio, L.(2019) J Biological Chem 294: 17117-17130
- PubMed: 31471321 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009223
- Primary Citation Related Structures:
6RS6, 6RS7, 6RS8, 6RS9 - PubMed Abstract:
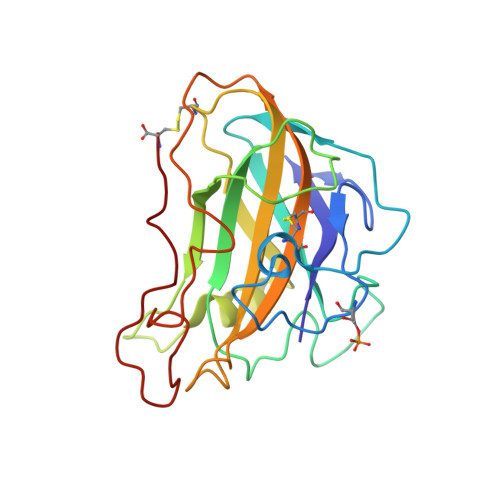
Lytic polysaccharide monooxygenases (LPMOs) are redox-enzymes involved in biomass degradation. All characterized LPMOs possess an active site of two highly conserved histidine residues coordinating a copper ion (the histidine brace), which are essential for LPMO activity. However, some protein sequences that belong to the AA9 LPMO family display a natural N-terminal His to Arg substitution (Arg-AA9). These are found almost entirely in the phylogenetic fungal class Agaricomycetes , associated with wood decay, but no function has been demonstrated for any Arg-AA9. Through bioinformatics, transcriptomic, and proteomic analyses we present data, which suggest that Arg-AA9 proteins could have a hitherto unidentified role in fungal degradation of lignocellulosic biomass in conjunction with other secreted fungal enzymes. We present the first structure of an Arg-AA9, Ls AA9B, a naturally occurring protein from Lentinus similis The Ls AA9B structure reveals gross changes in the region equivalent to the canonical LPMO copper-binding site, whereas features implicated in carbohydrate binding in AA9 LPMOs have been maintained. We obtained a structure of Ls AA9B with xylotetraose bound on the surface of the protein although with a considerably different binding mode compared with other AA9 complex structures. In addition, we have found indications of protein phosphorylation near the N-terminal Arg and the carbohydrate-binding site, for which the potential function is currently unknown. Our results are strong evidence that Arg-AA9s function markedly different from canonical AA9 LPMO, but nonetheless, may play a role in fungal conversion of lignocellulosic biomass.
- Department of Chemistry, University of Copenhagen, 2100 Copenhagen, Denmark.
Organizational Affiliation: