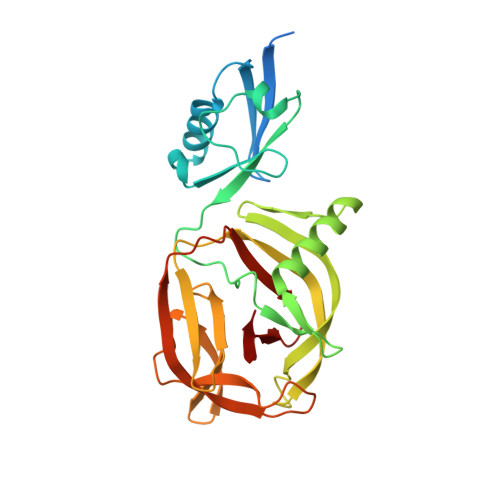
Crystal structures of CDC21-1 inteins from hyperthermophilic archaea reveal the selection mechanism for the highly conserved homing endonuclease insertion site.
Beyer, H.M., Mikula, K.M., Kudling, T.V., Iwai, H.(2019) Extremophiles 23: 669-679
- PubMed: 31363851 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00792-019-01117-4
- Primary Citation Related Structures:
6RPP, 6RPQ - PubMed Abstract:
Self-splicing inteins are mobile genetic elements invading host genes via nested homing endonuclease (HEN) domains. All HEN domains residing within inteins are inserted at a highly conserved insertion site. A purifying selection mechanism directing the location of the HEN insertion site has not yet been identified. In this work, we solved the three-dimensional crystal structures of two inteins inserted in the cell division control protein 21 of the hyperthermophilic archaea Pyrococcus abyssi and Pyrococcus horikoshii. A comparison between the structures provides the structural basis for the thermo-stabilization mechanism of inteins that have lost the HEN domain during evolution. The presence of an entire extein domain in the intein structure from Pyrococcus horikoshii suggests the selection mechanism for the highly conserved HEN insertion point.
- Research Program in Structural Biology and Biophysics, Institute of Biotechnology, University of Helsinki, P.O. Box 65, 00014, Helsinki, Finland.
Organizational Affiliation: