Self-assembled Pizza proteins
Noguchi, H., Voet, A.R.D.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 3fPizza6-SH | 256 | synthetic construct | Mutation(s): 0 | 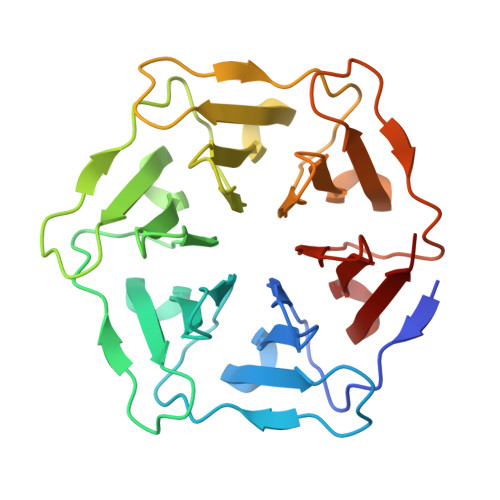 | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 41.954 | α = 90 |
| b = 53.264 | β = 102.39 |
| c = 43.452 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Research Foundation - Flanders | Belgium | G0E4717N, G0F9316N, G051917N |