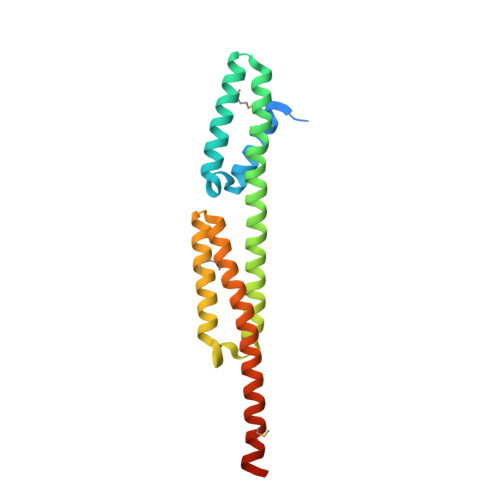
Crystal structure of the N-terminal domain of the major virulence factor BB0323 from the Lyme disease agent Borrelia burgdorferi.
Brangulis, K., Akopjana, I., Kazaks, A., Tars, K.(2019) Acta Crystallogr D Struct Biol 75: 825-830
- PubMed: 31478905 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319010751
- Primary Citation Related Structures:
6RJW, 6RJX - PubMed Abstract:
Lyme disease is an infection caused by the spirochete Borrelia burgdorferi after it is transmitted to a mammalian organism during a tick blood meal. B. burgdorferi encodes at least 140 lipoproteins located on the outer or inner membrane, thus facing the surroundings or the periplasmic space, respectively. However, most of the predicted lipoproteins are of unknown function, and only a few proteins are known to be essential for the persistence and virulence of the pathogen. One such protein is the periplasmic BB0323, which is indispensable for B. burgdorferi to cause Lyme disease and the function of which is associated with cell fission and outer membrane integrity. After expression and transport to the periplasm, BB0323 is cleaved into C-terminal and N-terminal domains by the periplasmic serine protease BB0104. The resulting N-terminal domain is sufficient to ensure the survival of B. burgdorferi throughout the mouse-tick infection cycle. The crystal structure of the N-terminal domain of BB0323 was determined at 2.35 Å resolution. The overall fold of the protein belongs to the spectrin superfamily, with the characteristic interconnected triple-helical bundles known as spectrin repeats that function as linkers between different cell components in other organisms. Overall, the reported three-dimensional structure of the N-terminal domain of BB0323 not only reveals the molecular details of a protein that is essential for B. burgdorferi membrane integrity, cell fission and infectivity, but also suggests that spectrin repeats in bacteria are not limited to the EzrA proteins.
- Latvian Biomedical Research and Study Centre, Ratsupites 1 k-1, Riga, LV-1067, Latvia.
Organizational Affiliation: